- Research article
- Open Access
- Published:
Alternative splicing and nonsense-mediated decay of circadian clock genes under environmental stress conditions inArabidopsis
BMC Plant Biologyvolume14, Article number:136(2014)
Abstract
Background
的circadian clock enables living organisms to anticipate recurring daily and seasonal fluctuations in their growth habitats and synchronize their biology to the environmental cycle. The plant circadian clock consists of multiple transcription-translation feedback loops that are entrained by environmental signals, such as light and temperature. In recent years, alternative splicing emerges as an important molecular mechanism that modulates the clock function in plants. Several clock genes are known to undergo alternative splicing in response to changes in environmental conditions, suggesting that the clock function is intimately associated with environmental responses via the alternative splicing of the clock genes. However, the alternative splicing events of the clock genes have not been studied at the molecular level.
Results
We systematically examined whether major clock genes undergo alternative splicing under various environmental conditions inArabidopsis. We also investigated the fates of the RNA splice variants of the clock genes. It was found that the clock genes, includingEARLY FLOWERING 3(ELF3) andZEITLUPE(ZTL) that have not been studied in terms of alternative splicing, undergo extensive alternative splicing through diverse modes of splicing events, such as intron retention, exon skipping, and selection of alternative 5′ splice site. Their alternative splicing patterns were differentially influenced by changes in photoperiod, temperature extremes, and salt stress. Notably, the RNA splice variants ofTIMING OF CAB EXPRESSION 1(TOC1) andELF3were degraded through the nonsense-mediated decay (NMD) pathway, whereas those of other clock genes were insensitive to NMD.
Conclusion
Taken together, our observations demonstrate that the major clock genes examined undergo extensive alternative splicing under various environmental conditions, suggesting that alternative splicing is a molecular scheme that underlies the linkage between the clock and environmental stress adaptation in plants. It is also envisioned that alternative splicing of the clock genes plays more complex roles than previously expected.
Background
的circadian clock is an endogenous time-keeping system that coordinates the physiology and behavior of a living organism to its environment [1]。In plants, the clock modulates rhythmic leaf movement, elongation rate of hypocotyls, roots, and stems, stomata aperture, stem circumnutation, and flower opening [1,2]。
Three major interlocked feedback loops constitute the plant circadian clock: the central loop, the morning loop, and the evening loop [3–5]。的central loop is mediated by the reciprocal repression between the morning-phased MYB transcription factors, CIRCADIAN CLOCK ASSOCIATED 1 (CCA1) and LATE ELONGATED HYPOCOTYL (LHY), and the evening-phased pseudo-response regulator TIMING OF CAB EXPRESSION 1 (TOC1) [6,7]。In the morning loop, CCA1 and LHY promote the transcription ofPSEUDO-RESPONSE REGULATOR 9(PRR9) andPRR7genes [8,9]。Closing the loop, the PRR members inhibit the transcription ofCCA1andLHYgenes by sequentially binding to the gene promoters from early morning (PRR9) through mid-day (PRR7) to evening (PRR5) [10,11]。的evening loop is illustrated by TOC1 and a hypothetical component Y, the expression of which is repressed by TOC1 and, in turn, activatesTOC1expression [12]。Recent studies have shown that three evening-phased factors, EARLY FLOWERING 3 (ELF3), ELF4, and LUX ARRHYTHMO (LUX), form the EVENING COMPLEX (EC), which repressesPRR9gene andLUXgene itself [13,14], indicating that the auto-inhibition of EC replaces the component Y in the evening loop [15]。
的circadian system is substantially influenced by external cues. Phytochrome- and cryptochrome-mediated light signals mediate the induction ofCCA1,LHY, andPRR9genes [8,16,17]。Temperatures also affect the amplitudes and rhythms of the clock gene expression [18]。In addition, growth hormones and abiotic stresses modulate the clock function. It has been observed that accumulation ofCCA1,TOC1, andGIGANTEA(GI) gene transcripts is differentially regulated by abscisic acid, brassinosteroid, and auxin [19]。High light stress inducesCCA1gene [20], linking the clock with plant stress adaptation.
的clock components are also regulated at the posttranscriptional and protein levels. It has been shown that the stability ofCCA1mRNA and the translation ofLHYmRNA are influenced by light [21,22]。In addition, the F-box protein ZEITLUPE (ZTL) is responsible for the dark-induced degradation of TOC1 protein [23]。Furthermore, temperature-dependent phosphorylation of CCA1 modulates its binding to target gene promoters [24]。Most recently, chromatin remodeling and alternative splicing of the clock genes have been described as fundamental processes in the regulation of the clock function [25]。
Some of the clock genes have been shown to undergo alternative splicing in response to abiotic stresses in plants [26,27], among which temperature regulation ofCCA1alternative splicing is best characterized.CCA1alternative splicing produces two protein isoforms, the full-size CCA1α form and the truncated CCA1β form that lacks the MYB DNA-binding motif [27]。CCA1β competitively inhibits CCA1α activity by forming nonfunctional heterodimers that are excluded from DNA binding.CCA1alternative splicing is suppressed by low temperatures. Under low temperature conditions, CCA1β production is reduced, and thus CCA1α activity is elevated, leading to the stimulation of freezing tolerance [27], linking the clock with temperature response.
Recently, it has been reported that alternatively spliced RNA isoforms of some clock genes are degraded through the nonsense-mediated decay (NMD) pathway [28–33], unlike the productive alternative splicing ofCCA1gene. NMD has evolved as an mRNA quality control mechanism that degrades mRNA molecules harboring premature termination codons (PTCs), which generate truncated proteins that are harmful to cellular energy metabolism, and those having aberrantly long 3′ untranslated regions (3′-UTRs) [32,33]。It is thus possible that alternative splicing serves as a precise mechanism for controlling the mRNA levels of the clock genes, depending on endogenous and external conditions.
In this study, we systematically investigated the alternative splicing patterns of major clock genes under various environmental conditions. We also examined the fates of the RNA splice variants. Our study shows that alternative splicing of the clock genes is differentially influenced by photoperiod and a variety of abiotic stresses. The results of our study show that although RNA splice variants of some clock genes are predicted to encode truncated versions of the authentic proteins, those of other clock genes do not appear to encode specific proteins and, instead, are degraded through the NMD pathway. It is envisioned that alternative splicing plays more complex roles in the clock function than previously expected.
Results
Major clock genes undergo extensive alternative splicing
On the basis of the prevalence of alternative splicing events in the plant circadian clock genes in the literature [20,26,27,34,35], we anticipated that alternative splicing of the core clock genes constitutes a critical component of the clock function. Previous reports have shown that alternative splicing ofCCA1is suppressed by low temperatures [20,27,35]。另一种蛋白质同种型(CCA1β)lacks the protein domain required for DNA binding, acts as a dominant negative regulator of the authentic CCA1 transcription factor (CCA1α), thus providing a self-regulatory circuit that links the clock with temperature stress response.
扩大我们的理解的功能relationship between the clock genes and environmental stress responses, we selected a group of major clock genes that constitutes the plant circadian clock and investigated whether these undergo alternative splicing and their alternative splicing patterns are altered under environmental stress conditions.
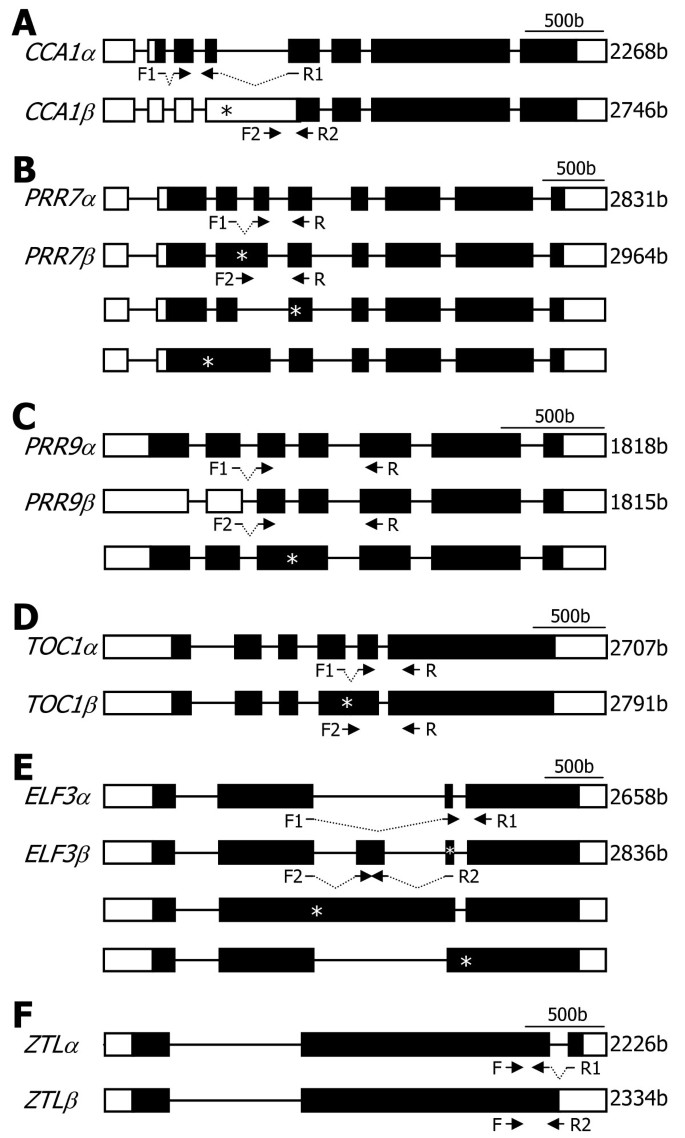
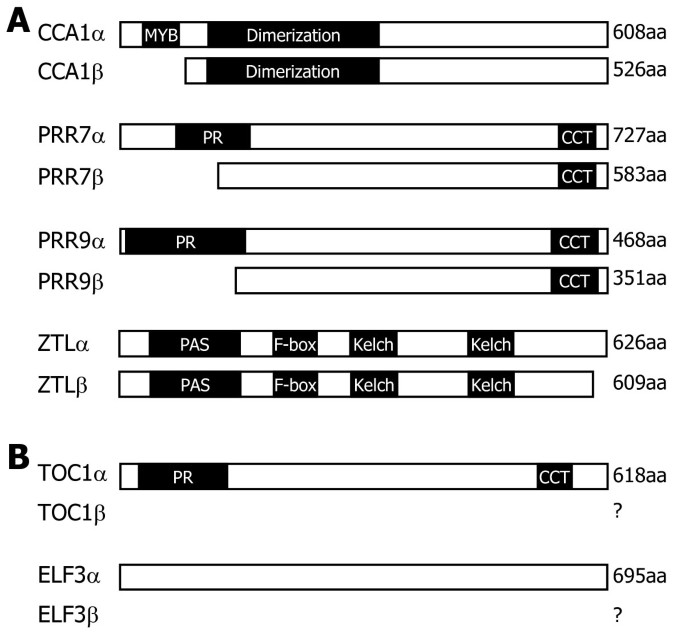
Analysis of gene structures deposited in the public databases and literature search revealed thatPRR7,PRR9,TOC1, andZTLgenes as well asCCA1gene undergo alternative splicing [26,27,34,35], each producing two or more RNA splice variants (Figure1). For each clock gene, theαtranscript represents the RNA splice variant that retains all the exons but do not have any introns. Theβtranscript represents the one that exists at the highest level among the RNA splice variants other than theαtranscript.
Genomic structures of major clock genes.的clock gene sequences were analyzed using the softwares provided by the TAIR database. The predicted genome structures ofCCA1(A),PRR7(B),PRR9(C),TOC1(D),ELF3(E), andZTL(F)genes are displayed. Black boxes depict exons. White boxes are 5′ and 3′ UTRs. F and R are primers used for RT-PCR analysis of the RNA splice variants (see Figure2). Theαtranscripts encode full-size, authentic proteins, and theβtranscripts encode truncated forms. Asterisks indicate premature termination codons (PTCs). b, bases. The genomic structure ofCCA1gene, which has already been reported by us [27], was included here for the benefit of the reader.
CCA1alternative splicing is mediated by the retention of intron 4 and introduces a PTC intoCCA1βtranscript (Figure1A).PRR7alternative splicing is somewhat complicated. It is mostly mediated by the retention of intron 3, resulting inPRR7βtranscript (Figure1B and Additional file1). A PTC is introduced into thePRR7βtranscript. Notably, it is also mediated by the skipping of exon 4 and the retention of introns 2 and 3 [26,34,35]。PRR9alternative splicing is unique, among others, in that the major alternatively spliced variant (PRR9β) is produced by selection of alternative 5′ splice site in intron 2 (Figure1C and Additional file2). The presence of two additional RNA splice variants has also been recently reported [26,34,35]。
A singleTOC1cDNA sequence was identified in the TAIR database. However, it has been shown that an alternative splicing event occurs by the retention of intron 4 [26,34], introducing a PTC intoTOC1βtranscript (Figure1D and Additional file3). It has been reported that RNA splice variants ofELF3gene are hardly detected in wild-type plants, but several RNA splice variants are detected in theskip-1μtant, which is defective in its splicing machinery [34], possibly due to the retention of intron 2 or 3 (Figure1E). We found that theELF3gene undergoes alternative splicing in wild-type plants (Additional file4). In addition, it was found that theELF3βtranscript is derived from the inclusion of a new alternative exon and a PTC is introduced into the splice variant.
的re are twoZTL-specific cDNA sequences (ZTLαandZTLβ) in the public database. Sequence comparison and direct sequencing of RT-PCR products revealed that theZTLalternative splicing is mediated by the retention of intron 2 (Figure1F and Additional file5). TheZTLβ-encoded protein has been considered as an authentic ZTL enzyme in the literature [23], which is probably because the abundance of theZTLβtranscript is much higher than that of theZTLαtranscript (see below).
的modes of splicing events are diverse in the clock genes
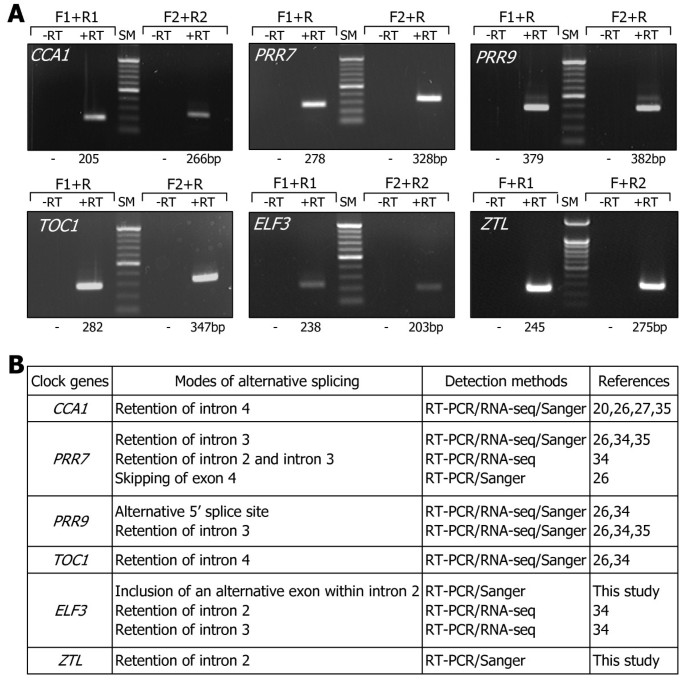
的abundances of RNA splice variants other thanαandβtranscripts were relatively very low in most cases ([26,34,35], this study). We therefore decided to further investigate only theαandβtranscripts for each clock gene. The predicted alternative splicing modes of the clock genes were verified by cloning of the RNA splice variants by RT-PCR and direct DNA sequencing (Additional files1,2,3,4, and5). Total RNA samples were subjected to RT-PCR using primer pairs that are specific to each RNA splice variant. The results showed that all the RT-PCR products have the sizes that were inferred from the predicted alternative splicing modes of the clock genes (Figure2A). No RT-PCR products were detected when reverse transcription was omitted prior to PCR amplifications, indicating that total RNA samples used were not contaminated with genomic DNA.
Detection of RNA splice variants of the clock genes. A. Detection of RNA splice variants by RT-PCR. Total RNA samples were isolated from 10-day-old Col-0 plants grown on MS-agar plates under LDs at peak ZT point for each clock gene and subject to RT-PCR. Gene-specific F and R primer sets, as indicated in Figure1, were used to detect the transcript isoforms of each clock gene. PCR reactions were also performed without reverse transcription (-RT) to verify the lack of genomic DNA contamination. The sizes of the PCR products are provided at the bottom of the figure. SM, DNA size marker. bp, base pair.B. Modes of splicing events. Detection methods for the alternative splicing events are listed in the 3rdcolumn with the references indicated in the 4thcolumn. The nucleotide sequences of the RNA splice variants were determined (This work) or verified by direct DNA sequencing in this work. RNA-seq, RNA sequencing. Sanger, DNA sequencing by Sanger method.
的modes of alternative splicing are diverse in the clock genes (Figure2B). Retention of specific introns mediates the alternative splicing ofCCA1,PRR7,TOC1,ZTL, andELF3genes. Exon skipping is involved inPRR7alternative splicing. Meanwhile, alternative 5′ splice site contributes toPRR9alternative splicing. Alternative splicing ofELF3gene was the most complicated. Retention of intron 2 or 3 has been implicated in theELF3alternative splicing [34]。However, direct sequencing of PCR products revealed that an additional RNA splice variant (ELF3β), which is probably most abundant among the splice variants, was produced by the inclusion of an alternative exon.
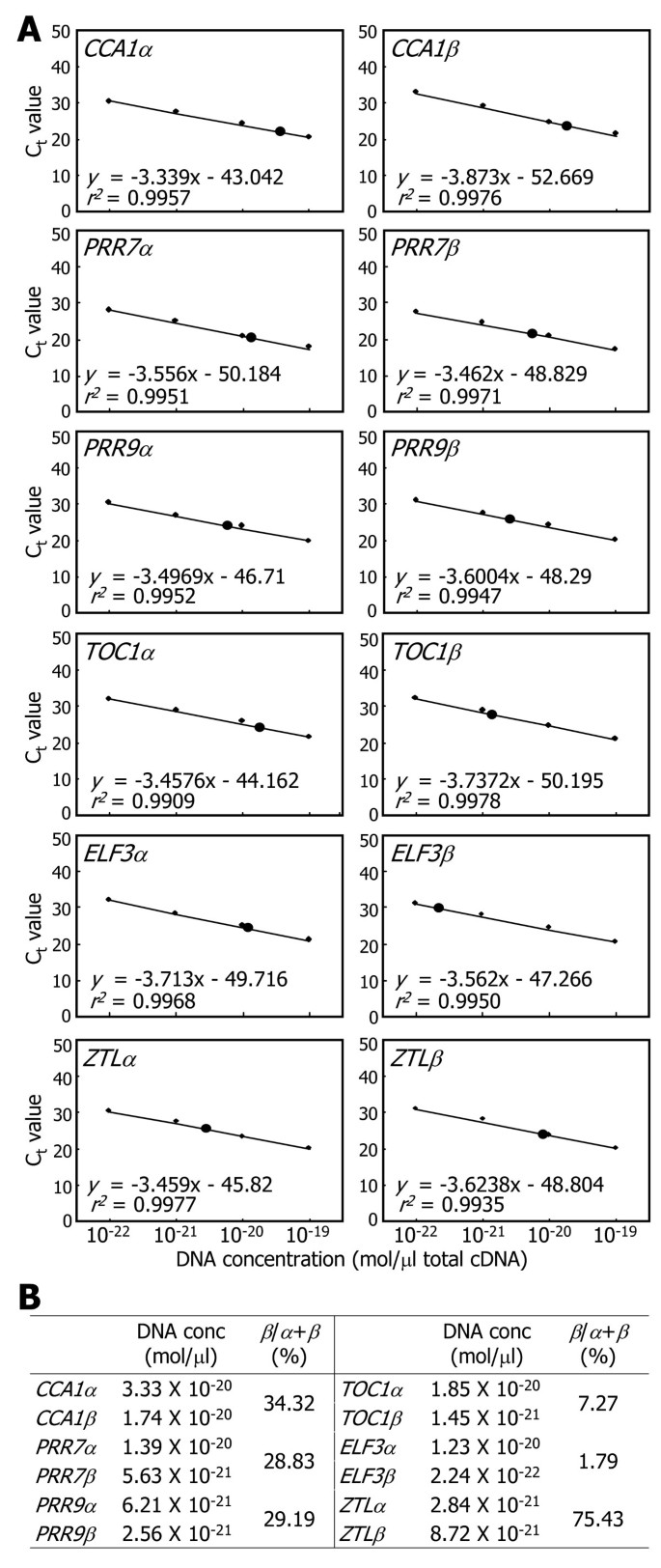
We measured the absolute amounts of the RNA splice variants of each clock gene by qRT-PCR analysis (Figure3A), as has been described previously [36,37]。Ten-day-old plants grown on MS-agar plates under long days (LDs, 16-h light and 8-h dark) were harvested at zeitgeber time (ZT) points of peak expression for individual clock genes (e.g. ZT0 forCCA1andZTL, ZT8 forPRR9, ZT4 forPRR7, and ZT12 forTOC1andELF3), thereby maximizing the detection sensitivity of a small quantity of mRNA. Absolute quantitation of the α and β RNA splice variants of each clock gene showed that the ratios (%) of β/α + β were variable among them (Figure3B). The ratio ofCCA1RNA splice variants was 34.32%, similar to what has been previously reported [27]。Those of the RNA splice variants ofPRR7andPRR9genes were approximately 29%. In contrast, those ofTOC1andELF3genes were relative low (<10%). One distinction wasZTLgene. Theβtranscript level was higher than that of theαtranscript (β/α + β = ~75%), which was in contrast to the other clock genes. It is assumed that the physiological significance of alternative splicing varies in each clock gene.
Absolute quantification of alternatively spliced RNA variants.Ten-day-old Col-0 plants grown on MS-agar plates were used for the extraction of total RNA samples. To maximize the sensitivity of detection, plants were harvested at the phase of peak expression for each gene. A series of 10-fold dilutions of plasmid DNA containing each gene sequence was used to generate a standard curve. The regression line from the dilution curve was used to determine the concentration of each RNA splice variant. Black circles represent the absolute amounts of RNA splice variants(A). CT, threshold cycle. The percentages ofβ/α + βwere calculated for each clock gene(B).
Some RNA splice variants of the clock genes are degraded by NMD
Alternatively spliced RNA variants containing a PTC enter either the productive or unproductive pathway. In the productive pathway, the mRNA is translated into a protein that is structurally distinct from the authentic protein. One example is the alternative splicing ofCCA1, in which the CCA1β protein isoform possesses protein domains required for dimer formation and transcriptional activation but lacks the MYB DNA-binding domain [27]。In contrast, in the unproductive pathway, the transcript is degraded via the NMD-mediated degradation pathway [30]。
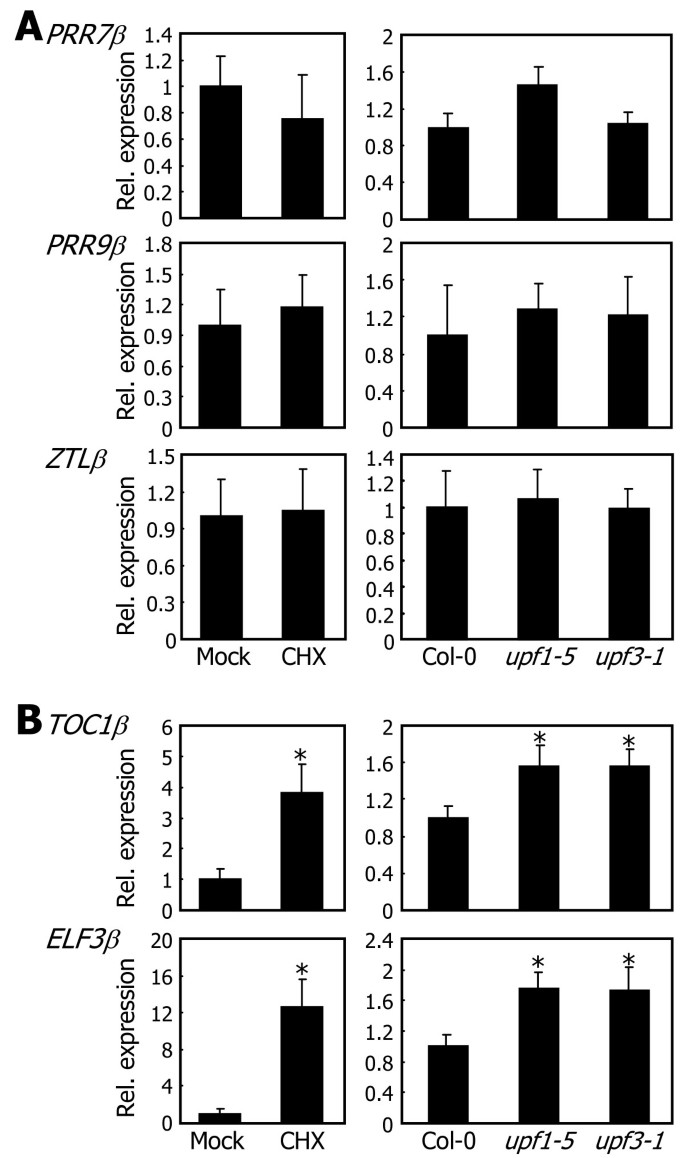
To investigate the fates of the RNA splice variants of the clock genes, we employed two assay systems: cycloheximide (CHX) treatment and NMD-defectiveArabidopsisμtants that are routinely employed for this purpose in the literature. NMD requires translation, and thus the translational inhibitor CHX suppresses the NMD-mediated degradation of mRNA [38,39]。Wild-typeArabidopsisplants were treated with CHX, and the levels ofβtranscripts were determined by qRT-PCR. It was found that whereas the levels ofPRR7β,PRR9β, andZTLβtranscripts were not influenced by CHX treatments (Figure4A, left panels), those ofTOC1βandELF3βtranscripts were significantly elevated after CHX treatments (Figure4B, left panels).
的fates of RNA splice variants.Steady-state levels of theβtranscripts were determined by qRT-PCR in Col-0 plants after CHX treatments (left panels) and in theupf1-5andupf3-1μtants (right panels). Biological triplicates were averaged and statistically treated using Studentt-test (*P < 0.01). Bars indicate standard error of the mean.A. The fates ofPRR7β,PRR9β, andZTLβtranscripts.B. The fates ofTOC1βandELF3βtranscripts.
We next examined theβtranscript levels of each clock gene in theupf1-5andupf3-1 Arabidopsisμtants, in which the NMD pathway is impaired [40,41]。的levels ofPRR7β,PRR9β, andZTLβtranscripts in the mutants were comparable to those in wild-type plants (Figure4A, right panels). In contrast, those ofTOC1βandELF3βtranscripts were higher by approximately two-fold in the mutants than in wild-type plants (Figure4B, right panels). We also examined the levels ofTOC1βandELF3βtranscripts in theupf1-5andupf3-1μtants under heat conditions. Theβtranscript levels were even higher in the mutants than in wild type plants when grown at 37°C (Additional file6). The more prominent differences in theTOC1βandELF3βtranscript levels at 37°C is due to the heat-induced alternative splicing ofTOC1andELF3genes (see below). Based on these observations, it was concluded that whereas thePRR7β,PRR9β, andZTLβtranscripts are likely to encode specific proteins, like theCCA1βtranscript [27], theTOC1βandELF3βtranscripts are probably targeted by NMD. The sensitivity of theTOC1βandELF3βtranscripts to NMD is also consistent with the notion that the steady-state levels of NMD target mRNAs were relatively low in many cases [30,42,43]。
Since the identities ofZTLαandZTLβtranscripts are currently unclear ([23], this work), we also examined the effects of CHX andupf1-5andupf3-1μtations on the accumulation ofZTLαtranscript. We found that theZTLαtranscript level was not affected by CHX treatments (Additional file7). It was also unaltered in theupf1-5andupf3-1μtants, like that ofZTLβtranscript under identical assay conditions. It is therefore likely that theZTLαtranscript is not targeted by NMD and, instead, encodes a distinct protein, like theZTLβtranscript.
组件defec蛋白亚型的时钟tive in different functional domains
Some RNA splice variants, such asPRR7β,PRR9β, andZTLβtranscripts that are insensitive to NMD, were predicted to encode truncated proteins that harbor altered protein structural organizations, as has been demonstrated with CCA1β protein isoform [27]。In many cases, these structural alterations in the truncated forms include deletions, insertions, or substitutions of certain protein domains [44]。
We analyzed the structural organization of the predicted protein isoforms of CCA1, PRR7, PRR9, and ZTL using the analysis tools in the SMART and Pfam databases (http://smart.embl-heidelberg.de/andhttp://pfam.sanger.ac.uk/, respectively). The amino acid sequences of the protein isoforms were obtained either from the TAIR database or deduced from the nucleotide sequences of RT-PCR products.
Two possible translation products were deduced fromPRR7βtranscript. One protein isoform would be a truncated form containing the N-terminal pseudo-receiver (PR) domain, which is generated by the translation from the start codon to PTC (Figure1). In this translation scheme, thePRR7βtranscript harbors a long 3′-UTR. It has been previously shown that alternatively spliced RNA variants having long 3′-UTRs are frequently targeted by NMD [30]。的other protein isoform is a truncated form lacking the N-terminal PR domain, which was marked as PRR7β (Figure5A). On the basis of the structural similarity of PRR7β to CCA1β and PRR9β and the insensitivity ofPRR7βtranscript to CHX, we believe that thePRR7βtranscript encodes the PRR7β protein that harbors the N-terminal truncation.
Domain structures of alternatively spliced protein isoforms.的protein domain structures were analyzed using the SMART and Pfam databases. Black boxes indicate the conserved protein domains. PR, pseudo-receiver; CCT, CONSTANS, CONSTANS-LIKE, and TOC1; PAS, Per-ARNT-Sim; aa, amino acid.A. Protein domain structures of CCA1, PRR7, PRR9, and ZTL and their protein isoforms.B. Protein domain structures of TOC1 and ELF3.
PRR7β and PRR9β protein isoforms possess the CONSTANS, CONSTANS-LIKE, and TOC1 (CCT) domains but lack the N-terminal PR domain (Figure5), which mediates interactions with other proteins, such as PRR5 [23,45–47]。的overall structures of the predicted ZTLα and ZTLβ protein isoforms were similar to each other except for the short C-terminal sequence. The ZTLβ isoform is slightly smaller than the ZTLα isoform by 17 residues (Figure5A). We were unable to identify any distinct protein motifs in the C-terminal region of the ZTL proteins, and thus it is currently unclear whether the two ZTL protein isoforms are functionally distinct or not.
Our data showed thatTOC1βandELF3βtranscripts were targeted by NMD and were not expected to encode any proteins (Figure5B).
Short days suppress the alternative splicing ofTOC1andELF3genes
Plants use the circadian clock to monitor daylength changes in inducing seasonal developmental responses [48,49]。We therefore hypothesized that photoperiod influences the alternative splicing patterns of the clock genes.
Arabidopsisplants were entrained to either LDs or short days (SDs, 8-h light and 16-h light), and the levels of alternatively spliced RNA variants were compared by qRT-PCR. The amplitudes and rhythms ofCCA1andPRR7gene expression were not detectably altered under SDs (Figure6A and B, respectively). Under SDs,PRR9gene was induced, but its expression rhythms were maintained (Figure6C). It seems that the alternative splicing of the morning-phased genes is not discernibly influenced by photoperiod.
Effects of photoperiod on the alternative splicing of the clock genes.Ten-day-old Col-0 plants grown on MS-agar plates under either LDs or SDs were harvested at the indicated ZT points for the extraction of total RNA samples. The levels of the RNA splice variants ofCCA1(A),PRR7(B),PRR9(C),TOC1(D),ELF3(E), andZTL(F)genes were determined by qRT-PCR. Biological triplicates were averaged. Bars indicate the standard error of the mean.
We observed that the levels ofTOC1αandELF3αtranscripts were higher under SDs than under LDs, evidently during the night (Figure6D and E, respectively). In contrast, the levels ofTOC1βtranscript were lower during the night under SDs, and those ofELF3βtranscript were not altered under identical conditions compared with LDs. Notably, the peak level of theTOC1βtranscript shifted from ZT12 under LDs to ZT8 under SDs. Together, these observations indicate that SDs suppress the alternative splicing of theTOC1andELF3genes. There were no discernible effects of SDs on the alternative splicing ofZTLgene (Figure6F).
Low temperatures suppressCCA1andELF3alternative splicing but induceTOC1alternative splicing
Low temperatures dampen the cyclic expression of the clock genes, resulting in the repression of the clock function [18]。Similarly, low temperatures suppress the alternative splicing ofCCA1, and the imbalance between CCA1α and CCA1β protein isoforms leads to disturbed circadian rhythms and induction of freezing tolerance [27]。It was therefore suspected that low temperatures would also affect the alternative splicing of other clock genes.
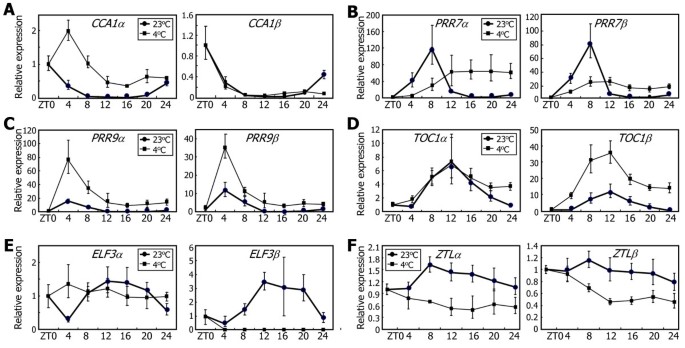
Arabidopsisplants were exposed to 4°C, and the levels of alternatively spliced RNA variants were measured by qRT-PCR. To eliminate the effects of light–dark transitions, the assays were conducted under continuous light conditions. It was found that the rhythmic accumulation patterns ofαandβtranscripts of the clock genes were significantly altered at low temperatures.CCA1alternative splicing was suppressed at low temperatures; the levels ofCCA1αtranscripts were elevated, whereas those ofCCA1βtranscripts remained low (Figure7A), as previously described [27]。
Effects of low temperatures on the alternative splicing of the clock genes.Ten-day-old Col-0 plants grown on MS-agar plates under LDs were transferred to 4°C under continuous light conditions. Whole plant materials were harvested at the indicated ZT points. The levels of the RNA splice variants ofCCA1(A),PRR7(B),PRR9(C),TOC1(D),ELF3(E), andZTL(F)genes were determined by qRT-PCR. Biological triplicates were averaged. Bars indicate the standard error of the mean.
的levels of bothPRR7αandPRR7βtranscripts were lower during the subjective day and higher during the subjective night compared to those at 23°C (Figure7B), indicating that low temperatures do not affectPRR7可变剪接但减少其节奏的经验ression. PRR9 is functionally redundant with PRR7 [12]。However, the effects of low temperatures onPRR9expression were distinct from those onPRR7expression. The levels of bothPRR9αandPRR9βtranscripts were markedly elevated at low temperatures throughout the time course, and the rhythmic expression was enhanced (Figure7C), indicating that low temperatures do not affect its alternative splicing.
的effects of low temperatures on the alternative splicing of the evening-phased genes were quite diverse. While the levels ofTOC1αtranscripts remained unchanged, those ofTOC1βtranscripts were markedly higher at low temperatures (Figure7D), indicating that low temperatures induceTOC1alternative splicing. The levels ofELF3αtranscripts were largely unaffected but loosed rhythmicity at low temperatures (Figure7E). In contrast, the levels ofELF3βtranscripts were drastically reduced, showing thatELF3alternative splicing is suppressed at low temperatures.ZTLexpression was suppressed at low temperatures, but its alternative splicing remained unaltered (Figure7F).
Heat induces the alternative splicing ofCCA1,PRR7,TOC1, andELF3genes
Heat stress has become an important issue in the field because of recent global warming that extensively affects plant growth and distribution [50]。Because the clock is entrained at least in part by temperature, heat would certainly influence the clock function. However, little is known about the relationship between heat stress and the clock. We therefore examined the effects of heat on the alternative splicing of the clock genes. The heat assays were performed under continuous light conditions to eliminate the effects of light–dark transitions.
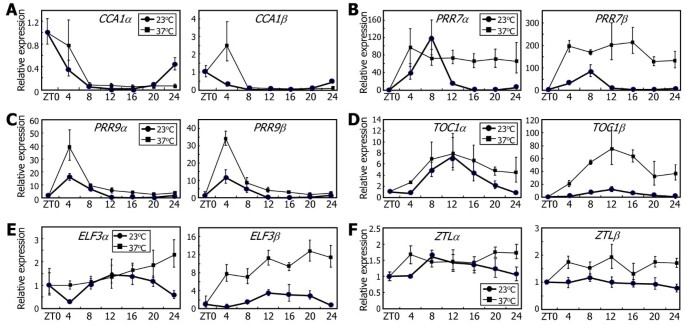
Interestingly, the balance betweenαandβtranscripts varied among different clock genes, whereas most clock genes were induced at 37°C. The levels ofCCA1βandPRR7βtranscripts were significantly elevated at some ZT points after heat treatments (Figure8A and B, respectively), showing that their alternative splicing was accordingly induced. The levels ofPRR9αandPRR9βtranscripts were elevated to a similar degree, showing that its alternative splicing is not affected by heat (Figure8C). The elevation ofTOC1βandELF3βtranscript levels were more prominent than that ofTOC1αandELF3αtranscript levels (Figure8D and E, respectively), suggesting that their alternative splicing was induced by heat. Heat effects were marginal onZTLexpression. The levels of bothZTLαandZTLβtranscripts were slightly elevated after heat treatments (Figure8F).
Effects of heat on the alternative splicing of the clock genes.Ten-day-old Col-0 plants grown on MS-agar plates under LDs were transferred to 37°C under continuous light conditions. Whole plant materials were harvested at the indicated ZT points. The levels of the RNA splice variants ofCCA1(A),PRR7(B),PRR9(C),TOC1(D),ELF3(E), andZTL(F)genes were determined by qRT-PCR. Biological triplicates were averaged. Bars indicate the standard error of the mean. Note that the expression data at 23°C are identical to those in Figure7.
High salinity suppressesELF3alternative splicing
Salt stress influences plant growth and developmental processes, such as flowering time, which is closely associated with the clock function [51–53]。We therefore examined the effects of high salinity on the alternative splicing of the clock genes.
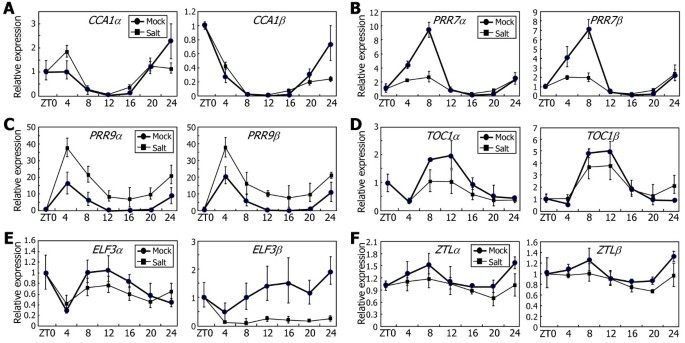
It appeared thatCCA1andZTLgenes are not influenced by high salinity (Figure9A and F, respectively). Notably,PRR7andTOC1genes were suppressed by high salinity. The levels of bothαandβtranscripts of these clock genes were reduced under high salt conditions (Figure9B and D, respectively), showing that their alternative splicing is not influenced by high salinity.ELF3gene was also suppressed by high salinity (Figure9E), but the reduction ofELF3βtranscript level was more prominent than that ofELF3αlevel, showing thatELF3alternative splicing is suppressed by high salinity. The levels ofPRR9αandPRR9βtranscripts increased with high salinity (Figure9C), indicating thatPRR9gene is induced but its alternative splicing is not affected by high salinity.
Effects of high salinity on the alternative splicing of the clock genes.Ten-day-old Col-0 plants grown on MS-agar plates under LDs were transferred to hydroponic MS medium containing 200 mM NaCl. The levels of the RNA splice variants ofCCA1(A),PRR7(B),PRR9(C),TOC1(D),ELF3(E), andZTL(F)genes were determined by qRT-PCR. Biological triplicates were averaged. Bars indicate the standard error of the mean.
Discussion
Effects of environmental conditions on the alternative splicing of the clock genes
Rhythmic expression of stress response genes and distinct phenotypes of clock mutants under abiotic stress conditions underscore the close connection between the circadian clock and environmental stress response in plants. One of the best-understood mechanisms is the clock control of C-REPEAT BINDING FACTOR(CBF) genes that play a pivotal role in cold stress response [18,54]。的central oscillators CCA1 and LHY regulate the expression of theCBFgenes by binding directly to their gene promoters [55]。的CBFgenes are also directly regulated by PRR9, PRR7, and PRR5 [56,57]。In addition, the transcription of CBF target genes, such asCOLD REGULATED 15 A(COR15A) andRESPONSIVE TO DISSECATION 29 A(RD29A) [58,59], is clock-controlled [18]。
Altered stress responses of various clock mutants further support the clock control of abiotic stress adaptation. Theprr9 prr7 prr5triple mutants exhibit enhanced resistance to drought and cold stresses [60]。TOC1-deficient mutants display drought-tolerant phenotypes [61]。In addition, it has been shown thatArabidopsisplants that are defective inCCA1,LHY, andGIgenes are susceptible to freezing temperatures [55,62]。
Although the linkage between the clock and environmental stress responses has been widely explored, molecular mechanisms and underlying signaling schemes have not been studied at the molecular level in most cases. It has been reported that low temperatures reduce the amplitude of the clock gene expression [18]。Meanwhile, it is known that the clock genes are regulated by extensive alternative splicing, which is influenced by low temperatures. It is therefore evident that alternative splicing of the clock genes should be taken into the interpretation of the expression analysis data under abiotic stress conditions.
This study shows that a group of major clock genes undergoes alternative splicing through a variety of splicing modes, such as intron retention, exon skipping, and selection of alternative 5′ splice sites, resulting in two or more RNA splice variants for each clock gene. It was also found that photoperiod and abiotic stresses, such as temperature extremes and high salinity, broadly affect the alternative splicing of the clock genes. On the basis of the effects of CHX on the relative levels of RNA splice variants and expression assays in NMD-defective mutants, we propose that the alternative splicing ofCCA1,PRR7,PRR9, andZTLgenes is productive with RNA splice variants encoding distinct proteins. In contrast, the RNA splice variants ofTOC1andELF3genes are predicted to be degraded through the NMD-mediated degradation pathway.
Collectively, our data strongly support the notion that the major clock genes are also regulated at the posttranscriptional level through alternative splicing in addition to the transcriptional control under both normal and environmental stress conditions. Alternative splicing-mediated control of the clock genes would serve as a molecular scheme that incorporates environmental stress signals into the clock, as has been verified withCCA1alternative splicing that links low temperature signals with the clock [27]。
In this work, we focused on two major RNA splice variants of each clock gene, although additional RNA splice variants have been identified or predicted for some of the clock genes examined (Figure1). More works on the additional RNA splice variants are required to further extend our understanding on the linkage between alternative splicing events of the clock genes and environmental stress responses. Searching for a full set of RNA splice variants of each clock gene, as has been performed by RNA sequencing method [20], will also be helpful for the elucidation of the clock function in abiotic stress adaptation. We are currently working on plants that are impaired in the alternative splicing of each clock gene and those expressing a specific RNA splice variant to investigate the physiological roles of the alternative splicing of the clock genes.
Function of alternative protein isoforms
Recent studies have shown that protein isoforms that lack specific functional domains, which are produced by the alternative splicing of transcription factor genes, act as competitive inhibitors of the authentic transcription factors by forming nonfunctional heterodimers [27,63]。最合适的机制是dominant negative regulation of the CCA1 transcription factor (CCA1α) by the protein isoform CCA1β. While the CCA1β isoform possesses protein domains required for dimmer formation and transcriptional activation, it lacks the MYB domain necessary for DNA binding [27]。的refore, CCA1β is capable of interacting with CCA1α, forming CCA1α-CCA1β heterodimers that are excluded from DNA binding.
According to the domain structures of the protein isoforms encoded by the NMD-insensitive RNA splice variants, the PRR7β and PRR9β protein isoforms are predicted to function in a way that is distinct from that of CCA1β. Unlike CCA1β that lacks the MYB DNA-binding domain, PRR7β and PRR9β have the CCT domain, which is responsible for DNA binding, but lack the PR domain that mediates protein-protein interactions [7,23,45–47]。A plausible working mechanism of the PRR7β and PRR9β protein isoforms would be that they compete with the authentic PRR7α and PRR9α transcription factors for binding to the promoters of target genes, as has been previously proposed [64]。Further investigations are required to determine the functional modes of PRR7β and PRR9β.
Two RNA splice variants ofZTL基因也对NMD,两个蛋白质isoforms, ZTLα and ZTLβ, are expected to be produced. The ZTLα and ZTLβ isoforms are identical except for the far C-terminal sequences; the former is larger than the latter by 17 residues. The functional mode of ZTLβ thus might differ from the β protein isoforms of other clock components. The lack of the C-terminal extension might also influence the protein conformation of the ZTLβ isoform, which would affect its substrate specificity or enzymatic activity. The smaller ZTLβ isoform has been annotated as the authentic ZTL protein in the literature [23,65]。We found that the level ofZTLβtranscript is much higher than that ofZTLαtranscript, which is in contrast to theα/βratios of other clock genes. It is currently unclear whether ZTLα or ZTLβ or both is an authentic enzyme. Phenotypic and physiological assays on transgenic plants that specifically express eitherZTLαorZTLβcDNA would help clarify this uncertainty.
NMD-mediated control of the clock gene expression
Unlike the NMD-insensitiveβtranscripts ofCCA1,PRR7,PRR9, andZTLgenes,TOC1βandELF3β成绩单显然是NMD的目标。的TOC1βandELF3βtranscripts possess sequence features that are frequently observed in NMD substrates, in which they have splice junctions downstream of the PTC and very long 3′-UTRs [30]。
Physiological roles of the NMD pathway are somewhat controversial. According to the “noise” hypothesis, NMD-sensitive RNA splice variants occur as a result of splicing error and are eventually removed through the NMD pathway [66]。In contrast, in the “regulated unproductive splicing and translation (RUST)” hypothesis, alternative splicing is coupled with NMD as a regulatory mechanism for monitoring the abundance of full-size RNA splice variants [67]。We believe that the RUST hypothesis fits well into the alternative splicing of theTOC1andELF3genes, based on the following reasons. First, the RUST hypothesis depicts that alternative splicing occurs through distinct modes of splicing events [26,34]。We found that the alternative splicing of theTOC1andELF3genes is mediated by the retention of specific introns, supporting the notion that their alternative splicing is a regulated process rather than a simple splicing error. Second, their alternative splicing is regulated by environmental factors in a discrete manner. Production of theELF3βtranscript is suppressed by cold and high salinity conditions but induced under heat stress conditions. Third, whereas their alternative splicing is markedly influenced by abiotic stresses, the levels ofTOC1αandELF3αtranscripts are less affected under identical conditions. However, it is still possible that the RNA splice variants may be at least in part translated into proteins. It has recently been reported that some NMD targets are stabilized and translated into proteins under certain conditions [68]。
Conclusions
We investigated the alternative splicing events of major clock genes under various environmental conditions and the sensitivity of their RNA splice variants to NMD. Alternative splicing patterns of the clock genes were differently affected by changes in photoperiod and abiotic stresses, such as cold, heat, and high salinity. Based on the results of this study, we propose that alternative splicing of the clock genes, either by producing truncated isoforms that act as self-regulators or by regulating the abundance of full-size transcripts at the posttranscriptional level, contributes to the precise regulation of the clock function, particularly under fluctuating environmental conditions. It may also serve as a web that integrates environmental stress signals into the clock, providing an adaptation strategy in response to unpredictable environmental changes.
Methods
Bioinformatics software
Gene sequences and their exon-intron structures were obtained from theArabidopsisInformation Resource (TAIR,http://www.arabidopsis.org/). Alternative splicing modes ofPRR7andTOC1genes, which have not been annotated in TAIR, were predicted based on the sequence analysis and the previous reports describing the predicted types of alternative splicing [26,34]。ForELF3gene that has not been studied, the presence of alternatively spliced RNA variants was verified by direct sequencing of RT-PCR products. Protein domain structures were predicted using the SMART (http://smart.embl-heidelberg.de/) and Pfam (http://www.sanger.ac.uk/Software/Pfam/) databases.
Plant materials and growth conditions
Arabidopsis thalianaecotype Columbia-0 (Col-0) was used, unless otherwise specified. Theupf1-5andupf3-1μtants, which have been previously described [26,34], were kindly provided by Dr. Jeong Sheop Shin (Korea University, Seoul, Korea) and Dr. Hee-Jeong Jeong (Kyung Hee University, Yongin, Korea). Plants were grown on ½ X Murashige & Skoog media containing 0.6% (w/v) agar (hereafter referred to as MS-agar plates) in a growth chamber set at 23°C with relative humidity of 60% under either long day conditions (LDs, 16-h light and 8-h dark) or short day conditions (SDs, 8-h light and 16-h dark) with white light illumination (120 μM photons m-2s-1) provided by fluorescent FLR40D/A tubes (Osram, Seoul, Korea).
Analysis of gene transcript levels
提取总RNA样本的适当的plant materials and RT-PCR conditions have been described previously [69]。Total RNA samples were extensively pretreated with an RNase-free DNase to eliminate contaminating genomic DNA prior to analysis.
Quantitative real-time RT-PCR (qRT-PCR) was employed to determine the levels of gene transcripts. RNA sample preparation, reverse transcription, and quantitative PCR were conducted according to the rules that have been proposed to ensure reproducible and accurate measurements [70]。
qRT-PCR reactions were performed in 96-well blocks using an Applied Biosystems 7500 Real-Time PCR System (Foster City, CA) using the SYBR Green I master mix in a volume of 20 μl. The PCR primers were designed using the Primer Express Software installed in the system and listed in Additional file8. The two-step thermal cycling profile used was 15 s at 94°C and 1 min at 68°C. TheeIF4Agene (At3g13920) was included in the reactions as internal control to normalize the variations in the amounts of cDNA used [71]。All qRT-PCR reactions were performed in biological triplicates using RNA samples extracted from three independent plant materials grown under identical conditions. The comparative ΔΔCTmethod was used to evaluate the relative quantities of each amplified product in the samples. The threshold cycle (CT) was automatically determined for each reaction using the default parameters of the system. The specificity of PCR reactions was determined by melt curve analysis of the amplified products using the standard methods installed in the system.
Absolute quantification of gene transcripts
Absolute quantification of gene transcripts was conducted as previously described [27]。的cDNAs of alternatively spliced RNA variants were subcloned into a pGADT7 vector (Clontech, Mountain View, CA), and the absolute standard curve of each transcript was obtained by a series of 10-fold dilutions covering from 10-19to 10-23mol/μl, as described elsewhere [36,37]。Quantitative RT-PCR was conducted using a SYBR Green I master mix (Applied Biosystems) with splice variant-specific primers listed in Additional file8.
Abiotic stress treatments
Arabidopsisplants grown for 10 days on MS-agar plates under LDs were used for abiotic stress treatments. For cold and heat treatments, plants were incubated at 4°C or at 37°C under continuous light conditions for appropriate time durations before harvesting plant materials. To examine the effects of high salinity on the alternative splicing of the clock genes, plants were transferred to MS liquid medium containing 200 mM NaCl under continuous light conditions for appropriate time durations.
Cycloheximide (CHX) treatments
的CHX treatments were performed as described elsewhere [29,30]。Ten-day-old plants grown on MS-agar plates were transferred to MS liquid medium containing 20 μM CHX. Following vacuum infiltration for 10 min, the plants were incubated for 5 h at 23°C under normal growth conditions before harvesting plant materials for total RNA extraction.
References
- 1.
Sanchez A, Shin J, Davis SJ: Abiotic stress and the plant circadian clock. Plant Signal Behav. 2011, 6: 223-231. 10.4161/psb.6.2.14893.
- 2.
Nakamichi N: Molecular mechanisms underlying theArabidopsiscircadian clock. Plant Cell Physiol. 2011, 52: 1709-1718. 10.1093/pcp/pcr118.
- 3.
McClung CR: Comes a time. Curr Opin Plant Biol. 2008, 11: 514-520. 10.1016/j.pbi.2008.06.010.
- 4.
Harmer SL: The circadian system in higher plants. Annu Rev Plant Biol. 2009, 60: 357-377. 10.1146/annurev.arplant.043008.092054.
- 5.
McClung CR: The genetics of plant clocks. Adv Genet. 2011, 74: 105-139.
- 6.
Alabadí D, Oyama T, Yanovsky MJ, Harmon FG, Más P, Kay SA: Reciprocal regulation between TOC1 and LHY/CCA1 within theArabidopsiscircadian clock. Science. 2011, 293: 880-883.
- 7.
Gendron JM, Pruneda-Paz JL, Doherty CJ, Gross AM, Kang SE, Kay SA:Arabidopsiscircadian clock protein, TOC1, is a DNA-binding transcription factor. Proc Natl Acad Sci USA. 2012, 109: 3167-3172. 10.1073/pnas.1200355109.
- 8.
Farré EM, Harmer SL, Harmon FG, Yanovsky MJ, Kay SA: Overlapping and distinct roles of PRR7 and PRR9 in theArabidopsiscircadian clock. Curr Biol. 2005, 15: 47-54. 10.1016/j.cub.2004.12.067.
- 9.
Harmer SL, Kay SA: Positive and negative factors confer phase-specific circadian regulation of transcription inArabidopsis. Plant Cell. 2005, 17: 1926-1940. 10.1105/tpc.105.033035.
- 10.
Nakamichi N, Kita M, Ito S, Yamashino T, Mizuno T: PSEUDO-RESPONSE REGULATORS, PRR9, PRR7 and PRR5, together play essential roles close to the circadian clock ofArabidopsis thaliana. Plant Cell Physiol. 2005, 46: 686-698. 10.1093/pcp/pci086.
- 11.
Nakamichi N, Kiba T, Henriques R, Mizuno T, Chua NH, Sakakibara H: PSEUDO-RESPONSE REGULATORS 9, 7, and 5 are transcriptional repressors in theArabidopsiscircadian clock. Plant Cell. 2010, 22: 594-605. 10.1105/tpc.109.072892.
- 12.
Locke JC, Kozma-Bognár L, Gould PD, Fehér B, Kevei E, Nagy F, Turner MS, Hall A, Millar AJ: Experimental validation of a predicted feedback loop in the multi-oscillator clock ofArabidopsis thaliana. Mol Syst Biol. 2006, 2: 59-
- 13.
Helfer A, Nusinow DA, Chow BY, Gehrke AR, Bulyk ML, Kay SA:LUX ARRHYTHMOencodes a nighttime repressor of circadian gene expression in theArabidopsiscore clock. Curr Biol. 2011, 21: 126-133. 10.1016/j.cub.2010.12.021.
- 14.
Nusinow DA, Helfer A, Hamilton EE, King JJ, Imaizumi T, Schultz TF, Farré EM, Kay SA: The ELF4-ELF3-LUX complex links the circadian clock to diurnal control of hypocotyl growth. Nature. 2011, 475: 398-402. 10.1038/nature10182.
- 15.
Pokhilko A, Fernández AP, Edwards KD, Southern MM, Halliday KJ, Millar AJ: The clock gene circuit inArabidopsisincludes a repressor with additional feedback loops. Mol Syst Biol. 2012, 8: 574-
- 16.
Somers DE, Devlin PF, Kay SA: Phytochromes and cryptochromes in the entrainment of theArabidopsiscircadian clock. Science. 1998, 282: 1488-1490.
- 17.
Martínez-García JF, Huq E, Quail PH: Direct targeting of light signals to a promoter element-bound transcription factor. Science. 2000, 288: 859-863. 10.1126/science.288.5467.859.
- 18.
Bieniawska Z, Espinoza C, Schlereth A, Sulpice R, Hincha DK, Hannah MA: Disruption of theArabidopsiscircadian clock is responsible for extensive variation in the cold-responsive transcriptome. Plant Physiol. 2008, 147: 263-279. 10.1104/pp.108.118059.
- 19.
Hanano S, Domagalska MA, Nagy F, Davis SJ: Multiple phytohormones influence distinct parameters of the plant circadian clock. Genes Cells. 2006, 11: 1381-1392. 10.1111/j.1365-2443.2006.01026.x.
- 20.
Filichkin SA, Priest HD, Givan SA, Shen R, Bryant DW, Fox SE, Wong WK, Mockler TC: Genome-wide mapping of alternative splicing inArabidopsis thaliana. Genome Res. 2010, 20: 45-58. 10.1101/gr.093302.109.
- 21.
Yakir E, Hilman D, Hassidim M, Green RM:CIRCADIAN CLOCK ASSOCIATED1transcript stability and the entrainment of the circadian clock inArabidopsis. Plant Physiol. 2007, 145: 925-932. 10.1104/pp.107.103812.
- 22.
Kim JY, Song HR, Taylor BL, Carré IA: Light-regulated translation mediates gated induction of theArabidopsisclock protein LHY. EMBO J. 2003, 22: 935-944. 10.1093/emboj/cdg075.
- 23.
Más P, Kim WY, Somers DE, Kay SA: Targeted degradation of TOC1 by ZTL modulates circadian function inArabidopsis thaliana. Nature. 2003, 426: 567-570. 10.1038/nature02163.
- 24.
Portoles年代,Mas P:功能起相互影响感情n protein kinase CK2 and CCA1 transcriptional activity is essential for clock temperature compensation inArabidopsis. PLoS Genet. 2010, 6: e1001201-10.1371/journal.pgen.1001201.
- 25.
Henriques R, Mas P: Chromatin remodeling and alternative splicing: pre- and post-transcriptional regulation of theArabidopsiscircadian clock. Semin Cell Dev Biol. 2013, 24: 399-406. 10.1016/j.semcdb.2013.02.009.
- 26.
James AB, Syed NH, Bordage S, Marshall J, Nimmo GA, Jenkins GI, Herzyk P, Brown JW, Nimmo HG: Alternative splicing mediates responses of theArabidopsis生物钟对温度变化. Plant Cell. 2012, 24: 961-981. 10.1105/tpc.111.093948.
- 27.
Seo PJ, Park MJ, Lim MH, Kim SG, Lee M, Baldwin IT, Park CM: A self-regulatory circuit of CIRCADIAN CLOCK-ASSOCIATED1 underlies the circadian clock regulation of temperature responses inArabidopsis. Plant Cell. 2012, 24: 2427-2442. 10.1105/tpc.112.098723.
- 28.
Shi C, Baldwin IT, Wu J:Arabidopsisplants having defects in nonsense-mediated mRNA decay factors UPF1, UPF2, and UPF3 show photoperiod-dependent phenotypes in development and stress responses. J Integr Plant Biol. 2012, 54: 99-114. 10.1111/j.1744-7909.2012.01093.x.
- 29.
Rayson S, Arciga-Reyes L, Wootton L, De Torres Zabala M, Truman W, Graham N, Grant M, Davies B: A role for nonsense-mediated mRNA decay in plants: pathogen responses are induced in Arabidopsis thaliana NMD mutants. PLoS One. 2012, 7: e31917-10.1371/journal.pone.0031917.
- 30.
Kalyna M, Simpson CG, Syed NH, Lewandowska D, Marquez Y, Kusenda B, Marshall J, Fuller J, Cardle L, McNicol J, Dinh HQ, Barta A, Brown JW: Alternative splicing and nonsense-mediated decay modulate expression of important regulatory genes inArabidopsis. Nucleic Acids Res. 2012, 40: 2454-2469. 10.1093/nar/gkr932.
- 31.
Syed NH, Kalyna M, Marquez Y, Barta A, Brown JW: Alternative splicing in plants-coming of age. Trends Plant Sci. 2012, 17: 616-623. 10.1016/j.tplants.2012.06.001.
- 32.
Drechsel G, Kahles A, Kesarwani AK, Stauffer E, Behr J, Drewe P, Rätsch G, Wachter A: Nonsense-mediated decay of alternative precursor mRNA splicing variants is a major determinant of theArabidopsissteady state transcriptome. Plant Cell. 2013, 25: 3726-3742. 10.1105/tpc.113.115485.
- 33.
Riehs-Kearnan N, Gloggnitzer J, Dekrout B, Jonak C, Riha K: Aberrant growth and lethality ofArabidopsisdeficient in nonsense-mediated RNA decay factors is caused by autoimmune-like response. Nucleic Acids Res. 2012, 40: 5615-5624. 10.1093/nar/gks195.
- 34.
Wang X, Wu F, Xie Q, Wang H, Wang Y, Yue Y, Gahura O, Ma S, Liu L, Cao Y, Jiao Y, Puta F, McClung CR, Xu X, Ma L: SKIP is a component of the spliceosome linking alternative splicing and the circadian clock inArabidopsis. Plant Cell. 2012, 24: 3278-3295. 10.1105/tpc.112.100081.
- 35.
Filichkin SA, Mockler TC: Unproductive alternative splicing and nonsense mRNAs: a widespread phenomenon among plant circadian clock genes. Biol Direct. 2012, 7: 20-10.1186/1745-6150-7-20.
- 36.
Bustin SA: Absolute quantification of mRNA using real-time reverse transcription polymerase chain reaction assays. J Mol Endocrinol. 2000, 25: 169-193. 10.1677/jme.0.0250169.
- 37.
Whelan JA, Russell NB, Whelan MA: A method for the absolute quantification of cDNA using real-time PCR. J Immunol Methods. 2003, 278: 261-269. 10.1016/S0022-1759(03)00223-0.
- 38.
Ishigaki Y, Li X, Serin G, Maquat LE: Evidence for a pioneer round of mRNA translation: mRNAs subject to nonsense-mediated decay in mammalian cells are bound by CBP80 and CBP20. Cell. 2001, 106: 607-617. 10.1016/S0092-8674(01)00475-5.
- 39.
Brogna S, Wen J: Nonsense-mediated mRNA decay (NMD) mechanisms. Nat Struct Mol Biol. 2009, 16: 107-113. 10.1038/nsmb.1550.
- 40.
Arciga-Reyes L, Wootton L, Kieffer M, Davies B: UPF1 is required for nonsense-mediated mRNA decay (NMD) and RNAi inArabidopsis. Plant J. 2006, 47: 480-489. 10.1111/j.1365-313X.2006.02802.x.
- 41.
Hori K, Watanabe Y: UPF3 suppresses aberrant spliced mRNA inArabidopsis. Plant J. 2005, 43: 530-540. 10.1111/j.1365-313X.2005.02473.x.
- 42.
Stauffer E, Westermann A, Wagner G, Wachter A: Polypyrimidine tract-binding protein homologues fromArabidopsisunderlie regulatory circuits based on alternative splicing and downstream control. Plant J. 2010, 64: 243-255. 10.1111/j.1365-313X.2010.04321.x.
- 43.
McGlincy NJ, Smith CW: Alternative splicing resulting in nonsense-mediated mRNA decay: what is the meaning of nonsense?. Trends Biochem Sci. 2008, 33: 385-393. 10.1016/j.tibs.2008.06.001.
- 44.
Wang P, Yan B, Guo JT, Hicks C, Xu Y: Structural genomics analysis of alternative splicing and application to isoform structure modeling. Proc Natl Acad Sci USA. 2005, 102: 18920-18925. 10.1073/pnas.0506770102.
- 45.
Wenkel S, Turck F, Singer K, Gissot L, Le Gourrierec J, Samach A, Coupland G: CONSTANS and the CCAAT box binding complex share a functionally important domain and interact to regulate flowering ofArabidopsis. Plant Cell. 2006, 18: 2971-2984. 10.1105/tpc.106.043299.
- 46.
Wang L, Fujiwara S, Somers DE: PRR5 regulates phosphorylation, nuclear import and subnuclear localization of TOC1 in theArabidopsiscircadian clock. EMBO J. 2010, 29: 1903-1915. 10.1038/emboj.2010.76.
- 47.
Para A, Farré EM, Imaizumi T, Pruneda-Paz JL, Harmon FG, Kay SA: PRR3 Is a vascular regulator of TOC1 stability in theArabidopsiscircadian clock. Plant Cell. 2007, 19: 3462-3473. 10.1105/tpc.107.054775.
- 48.
Millar AJ: Input signals to the plant circadian clock. J Exp Bot. 2004, 55: 277-283.
- 49.
Imaizumi T:Arabidopsiscircadian clock and photoperiodism: time to think about location. Curr Opin Plant Biol. 2010, 13: 83-89. 10.1016/j.pbi.2009.09.007.
- 50.
Long SP, Ort DR: More than taking the heat: crops and global change. Curr Opin Plant Biol. 2010, 13: 241-248.
- 51.
Zeller G, Henz SR, Widmer CK, Sachsenberg T, Rätsch G, Weigel D, Laubinger S: Stress-induced changes in theArabidopsis thalianatranscriptome analyzed using whole-genome tiling arrays. Plant J. 2009, 58: 1068-1082. 10.1111/j.1365-313X.2009.03835.x.
- 52.
Chan Z, Loescher W, Grumet R: Transcriptional variation in response to salt stress in commonly usedArabidopsis thalianaaccessions. Plant Physiol Biochem. 2013, 73: 189-201.
- 53.
Munns R, Tester M: Mechanisms of salinity tolerance. Annu Rev Plant Biol. 2008, 59: 651-681. 10.1146/annurev.arplant.59.032607.092911.
- 54.
Fowler SG, Cook D, Thomashow MF: Low temperature induction ofArabidopsis CBF1, 2, and3is gated by the circadian clock. Plant Physiol. 2005, 137: 961-968. 10.1104/pp.104.058354.
- 55.
Dong MA, Farré EM, Thomashow MF: Circadian clock-associated 1 and late elongated hypocotyl regulate expression of the C-repeat binding factor (CBF) pathway inArabidopsis. Proc Natl Acad Sci USA. 2011, 108: 7241-7246. 10.1073/pnas.1103741108.
- 56.
Nakamichi N, Kiba T, Kamioka M, Suzuki T, Yamashino T, Higashiyama T, Sakakibara H, Mizuno T: Transcriptional repressor PRR5 directly regulates clock-output pathways. Proc Natl Acad Sci USA. 2012, 109: 17123-17128. 10.1073/pnas.1205156109.
- 57.
Liu T, Carlsson J, Takeuchi T, Newton L, Farré EM: Direct regulation of abiotic responses by the Arabidopsis circadian clock component PRR7. Plant J. 2013, 76: 101-114.
- 58.
Shinozaki K, Yamaguchi-Shinozaki K: Molecular responses to dehydration and low temperature: differences and cross-talk between two stress signaling pathways. Curr Opin Plant Biol. 2000, 3: 217-223. 10.1016/S1369-5266(00)80068-0.
- 59.
Msanne J, Lin J, Stone JM, Awada T: Characterization of abiotic stress-responsiveArabidopsis thaliana RD29AandRD29Bgenes and evaluation of transgenes. Planta. 2011, 234: 97-107. 10.1007/s00425-011-1387-y.
- 60.
Nakamichi N, Kusano M, Fukushima A, Kita M, Ito S, Yamashino T, Saito K, Sakakibara H, Mizuno T: Transcript profiling of anArabidopsis PSEUDO RESPONSE REGULATORarrhythmic triple mutant reveals a role for the circadian clock in cold stress response. Plant Cell Physiol. 2009, 50: 447-462. 10.1093/pcp/pcp004.
- 61.
Legnaioli T, Cuevas J, Mas P: TOC1 functions as a molecular switch connecting the circadian clock with plant responses to drought. EMBO J. 2009, 28: 3745-3757. 10.1038/emboj.2009.297.
- 62.
Cao S, Ye M, Jiang S: Involvement ofGIGANTEAgene in the regulation of the cold stress response inArabidopsis. Plant Cell Rep. 2005, 24: 683-690. 10.1007/s00299-005-0061-x.
- 63.
Seo PJ, Kim MJ, Ryu JY, Jeong EY, Park CM: Two splice variants of the IDD14 transcription factor competitively form nonfunctional heterodimers which may regulate starch metabolism. Nat Commun. 2011, 2: 303-
- 64.
Seo PJ, Hong SY, Kim SG, Park CM: Competitive inhibition of transcription factors by small interfering peptides. Trends Plant Sci. 2011, 16: 541-549. 10.1016/j.tplants.2011.06.001.
- 65.
Somers DE, Schultz TF, Milnamow M, Kay SA:ZEITLUPEencodes a novel clock-associated PAS protein fromArabidopsis. Cell. 2000, 101: 319-329. 10.1016/S0092-8674(00)80841-7.
- 66.
Saltzman AL, Kim YK, Pan Q, Fagnani MM, Maquat LE, Blencowe BJ: Regulation of multiple core spliceosomal proteins by alternative splicing-coupled nonsense-mediated mRNA decay. Mol Cell Biol. 2008, 28: 4320-4330. 10.1128/MCB.00361-08.
- 67.
孟Lareau低频,布鲁克斯,Soergel哒,Q,布伦纳SE: The coupling of alternative splicing and nonsense-mediated mRNA decay. Adv Exp Med Biol. 2007, 623: 190-211. 10.1007/978-0-387-77374-2_12.
- 68.
Colak D, Ji SJ, Porse BT, Jaffrey SR: Regulation of axon guidance by compartmentalized nonsense-mediated mRNA decay. Cell. 2013, 153: 1252-1265. 10.1016/j.cell.2013.04.056.
- 69.
Kim YS, Kim SG, Lee M, Lee I, Park HY, Seo PJ, Jung JH, Kwon EJ, Suh SW, Paek KH, Park CM: HD-ZIP III activity is modulated by competitive inhibitors via a feedback loop inArabidopsisshoot apical meristem development. Plant Cell. 2008, 20: 920-933. 10.1105/tpc.107.057448.
- 70.
Udvardi MK, Czechowski T, Scheible WR: Eleven golden rules of quantitative RT-PCR. Plant Cell. 2008, 20: 1736-1737. 10.1105/tpc.108.061143.
- 71.
Gutierrez L, Mauriat M, Guénin S, Pelloux J, Lefebvre JF, Louvet R, Rusterucci C, Moritz T, Guerineau F, Bellini C, Van Wuytswinkel O: The lack of a systematic validation of reference genes: a serious pitfall undervalued in reverse transcription-polymerase chain reaction (RT-PCR) analysis in plants. Plant Biotechnol J. 2008, 6: 609-618. 10.1111/j.1467-7652.2008.00346.x.
Acknowledgements
We thank Drs. Jeong Sheop Shin and Hee-Jeong Jeong for kindly providing theupf1andupf3μtants. This work was supported by the International Exchange Program for University Researchers through the National Research Foundation of Korea (013-2011-1-C00048) and the Next-Generation BioGreen 21 Program (Plant Molecular Breeding Center No. 201203013055290010200) provided by the Rural Development Administration, Korea. It was also supported in part by the Human Frontier Science Program (RGP0002/2012).
Author information
Affiliations
Corresponding author
Additional information
Competing interests
的authors declare that they have no competing interests.
Authors’ contributions
CMP和YJK概念化项目和分析d the data. CMP, YJK, and MJP wrote the manuscript. YJK and MJP carried out the molecular assays on the alternative splicing of the clock genes. YJK, SGK, and ITB predicted the alternative splicing patterns of the clock genes. All authors discussed the results and approved the final form of the manuscript.
Young-Ju Kwon, Mi-Jeong Park contributed equally to this work.
Electronic supplementary material
Nucleotide sequence comparison of
Additional file 1:PRR7gDNA andPRR7βcDNA.的nucleotide sequence ofPRR7βcDNA was determined by DNA sequencing of RT-PCR product and aligned withPRR7genomic DNA (PRR7gDNA) using the ClustalW software (http://www.ebi.ac.uk/tools/msa/clustalw2/). Part of the aligned sequences containing exons 1, 2, 3, and 4 and introns 2, 3, and 4 was displayed. Intron 3, which is retained in thePRR7βtranscript as a result of alternative splicing, is underlined (blue). A PTC (premature termination codon) is introduced into thePRR7βtranscript (red asterisk). (PDF 65 KB)
Nucleotide sequence comparison of
Additional file 2:PRR9gDNA andPRR9βcDNA.的nucleotide sequence ofPRR9βcDNA was determined by DNA sequencing of RT-PCR product and aligned withPRR9使用ClustalW gDNA软件。讽刺笑星阿里•g的一部分ned sequences containing exons 1, 2, 3, 4, and 5 and introns 1, 2, 3, and 4 was displayed. The alternative splice site, which is used to produce thePRR9βtranscript, is underlined. Sequence analysis revealed that thePRR9βtranscript occurs by the alternative 5′ splice site in intron 2 (blue). (PDF 65 KB)
Nucleotide sequence comparison of
Additional file 3:TOC1gDNA andTOC1βcDNA.的nucleotide sequence ofTOC1βcDNA was determined by DNA sequencing of RT-PCR product and aligned withTOC1使用ClustalW gDNA软件。讽刺笑星阿里•g的一部分ned sequences containing exons 4, 5, and 6 and introns 3, 4, and 5 was displayed. The retained intron 4, which is included in theTOC1βtranscript as a result of alternative splicing, is underlined (blue). A PTC is introduced into theTOC1βtranscript (red asterisk). (PDF 86 KB)
Nucleotide sequence comparison of
Additional file 4:ELF3gDNA andELF3βcDNA.的nucleotide sequence ofELF3βcDNA was determined by DNA sequencing of RT-PCR product and aligned withELF3使用ClustalW gDNA软件。讽刺笑星阿里•g的一部分ned sequences containing exons 2 and 3 and intron 2 was displayed. The alternative exon within intron 2, which is included in theELF3βtranscript as a result of alternative splicing, is underlined. Red boxes indicate conserved ‘T’ and 'AG’ sequences at the 5′ and 3′ ends of introns. Sequence analysis revealed that theELF3βtranscript occurs by the inclusion of an alternative exon consisting of 178 nucleotides within intron 2 (blue). A PTC is introduced into theELF3βtranscript (red asterisk). (PDF 89 KB)
Nucleotide sequence comparison of
Additional file 5:ZTLgDNA andZTLαandZTLβcDNAs.的nucleotide sequences ofZTLαandZTLβcDNAs were determined by DNA sequencing of RT-PCR products and aligned withZTL使用ClustalW gDNA软件。讽刺笑星阿里•g的一部分ned sequences containing exons 2 and 3 and intron 2 was displayed. Intron 2, which is retained in theZTLβtranscript as a result of alternative splicing, is underlined (blue). A PTC is introduced into theZTLβtranscript (red asterisk). The 3′ untranslated region of theZTLβtranscript is shown in gray. (PDF 79 KB)
的fate of
Additional file 6:TOC1βandELF3βtranscripts under heat stress conditions.Ten-day-old Col-0 plants andupf1-5andupf3-1μtants grown on ½ X Murashige & Skoog media containing 0.6% (w/v) agar plates (hereafter referred to as MS-agar plates) were transferred to 37°C for 12 h before harvesting whole plant materials for the extraction of total RNA. Levels ofTOC1βandELF3βtranscripts were determined by quantitative real-time RT-PCR (qRT-PCR). Biological triplicates were averaged and statistically treated using Studentt-test (*P<0.01). Bars indicate standard error of the mean. (PDF 39 KB)
的fate of
Additional file 7:ZTLαtranscript.Plants were grown on MS-agar plates for 10 days under normal growth conditions. Ten-day-old Col-0 plants were transferred to liquid MS culture containing 20 μM cycloheximide (CHX). Following vacuum infiltration for 10 min, the plants were incubated for 5 h at 23°C under normal growth conditions before harvesting whole plant materials for the extraction of total RNA (left panel). Theupf1-5andupf3-1μtants were not treated with CHX (right panel). Levels ofZTLαtranscript were determined by qRT-PCR Biological triplicates were averaged. Bars indicate standard error of the mean. (PDF 39 KB)
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
Open AccessThis article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made.
的images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
To view a copy of this licence, visithttps://creativecommons.org/licenses/by/4.0/.
的Creative Commons Public Domain Dedication waiver (https://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Kwon, YJ., Park, MJ., Kim, SG.et al.Alternative splicing and nonsense-mediated decay of circadian clock genes under environmental stress conditions inArabidopsis.BMC Plant Biol14,136 (2014). https://doi.org/10.1186/1471-2229-14-136
Received:
Accepted:
Published:
Keywords
- Arabidopsis thaliana
- Circadian clock
- Transcription factor
- Alternative splicing
- Nonsense-mediated decay (NMD)
- Environmental stress