- Research article
- Open Access
- Published:
Identification and functional characterization of the chloride channel gene,GsCLC-c2from wild soybean
BMC Plant Biologyvolume19, Article number:121(2019)
Abstract
Background
The anionic toxicity of plants under salt stress is mainly caused by chloride (Cl−). Thus Cl−influx, transport and their regulatory mechanisms should be one of the most important aspects of plant salt tolerance studies, but are often sidelined by the focus on sodium (Na+) toxicity and its associated adaptations. Plant chloride channels (CLCs) are transport proteins for anions including Cl−and nitrate (NO3−), and are critical for nutrition uptake and transport, adjustment of cellular turgor, stomatal movement, signal transduction, and Cl−and NO3−homeostasis under salt stress.
Results
Among the eight soybeanCLCgenes, the tonoplast-localizedc2has uniquely different transcriptional patterns between cultivated soybean N23674 and wild soybean BB52. Using soybean hairy root transformation, we found thatGsCLC-c2over-expression contributed to Cl−and NO3−homeostasis, and therefore conferred salt tolerance, through increasing the accumulation of Cl−in the roots, thereby reducing their transportation to the shoots where most of the cellular damages occur. Also, by keeping relatively high levels of NO3−in the aerial part of the plant,GsCLC-c2could reduce the Cl−/NO3−ratio. Wild type GsCLC-c2, but not its mutants (S184P,E227VandE294G) with mutations in the conserved domains, is able to complementSaccharomyces cerevisiae△gef1Cl−sensitive phenotype. Using two-electrode voltage clamp onXenopus laevisoocytes injected withGsCLC-c2cRNA, we found that GsCLC-c2 transports both Cl−and NO3−with slightly different affinity, and the affinity toward Cl−was pH-independent.
Conclusion
This study revealed that the expression ofGsCLC-c2is induced by NaCl-stress in the root of wild soybean. The tonoplast localized GsCLC-c2 transports Cl−with a higher affinity than NO3−in a pH-independent fashion. GsCLC-c2 probably alleviates salt stressin plantathrough the sequestration of excess Cl−into the vacuoles of root cells and thus preventing Cl−from entering the shoots where it could result in cellular damages.
Background
Salinization of irrigation water and soil is one of the most serious environmental factors limiting crop productivity worldwide [1,2,3]。Sodium chloride (NaCl) is the most common salt found in saline soil. It hinders plant growth not only through ionic toxicity but also by causing water deficit inside the plant, i.e. osmotic stress, and imbalance or deficiency of other ions such as potassium (K+) and nitrate (NO3−) [4,5,6]。Plants developed different strategies to combat salt stress, including exclusion or compartmentalization of ions to reduce cytosolic Na+and Cl−, especially in the aerial parts [7,8]。Generally, crops such as cotton, rice and barley are more sensitive to Na+than Cl−, whereas tobacco, grape, potato, citrus and cultivated soybean, in which salt stress effects are mainly caused by Cl−, are known as “Cl−-sensitive” or “Cl−讨厌“植物8,9,10,11,12,13,14,15]。So far, researches on the physiological and molecular mechanisms of salt tolerance in plants have mostly focused on Na+有毒ity and adaptations. However, until recently, the salt injury caused by Cl−has been largely ignored [11,14]。
Cl−, as one of the essential micronutrient elements, is one of the main anions in plant cells besides NO3−, and is normally accumulated to macronutrient levels for enhancing plant growth and development. Cl−is involved in photosynthesis by stabilizing the water splitting system or oxygen-evolving complex of photosystem II (PSII), and also in stomatal movement, cellular osmotic pressure maintenance, electrical charge balance, turgor and pH regulation, nitrogen use efficiency, water-holding capacity and disease resistance [5,14,15,16]。Chloride deficiency leads to reduced plant growth and symptoms such as leaf chlorosis [15]。Despite its beneficial roles in plant nutrition, Cl−can be a major toxic element in the cytosol when plants are grown in salt-affected soils, where Cl−is predominant and can accumulate in excess in the shoot/leaf, adversely affecting plant growth [6,15]。During salt stress, the effects of Cl−can be additive or synergistic to those of Na+[11]。Excessive amounts of Cl−can lead to decreased uptake of NO3−, which is the most important nitrogen source for plants and the main monovalent anion in plant tissues and cells next to Cl−[14,17,18]。Both NO3−and Cl−are monovalent anions with similar ionic radii, and perform a similar role in maintaining cellular charge balance and turgor, and can often be transported by the same proteins [11]。To combat Cl−/salt stress in plants, an increased NO3−:Cl−ratio in the tissues or organs, similar to the well-described increased K+:Na+ratio in shoots, can play a positive role [9,14,19,20]。Although chloride is a beneficial micronutrient in its own right and not simply a “cheap osmoticum” for plants [15], its impacts on plants under salt stress are problematic [11]。The nature of Cl−有毒ity or the mechanisms of Cl−transport and detoxification during salt stress is far from being understood to such degrees as those of Na+[14,21]。Despite the general lack of information, a few reports have suggested that the control of Cl−transport from roots to shoots or the ability to maintain a low Cl−Cl的拍摄水平是关键决定因素−/salt tolerance in plants [8,10,12]。In the past few years, effects of Cl−有毒ity and functions of some essential Cl−transporters, such as the chloride channel proteins (CLCs), that mediate Cl−transport and homeostasis in plants under salt stress are drawing increasing research interests [11,15]。
Currently,CLCgenes have been identified in the genome of higher plants includingArabidopsis, tobacco, rice, potato, citrus, soybean, maize, and poplar with 7–8 gene members in each species (summarized in [21,22]). In total, 217 homologous CLC protein or gene sequences belonging to 34 species of plants have been retrieved from the Pfam database (http://pfam.xfam.org/), Uniprot database (http://www.uniprot.org/), and NCBI database [22,23]。The large number of plant CLC proteins, with their dual functions of NO3−and Cl−transport, and the differences in their respective affinity for NO3−or Cl−pose a degree of technical difficulties and uncertainties in the results from in-depth researches [24,25,26]。
The genetic diversity between cultivated crops and their wild relatives provides rich resources for trait and gene discovery that have yet to be sufficiently utilized. The cultivated soybean (G. max) is the most important legume crop in the world, offering high-quality protein and oil for human food and animal feed [27]。When exposed to NaCl stress, Cl−in soybean plants is usually accumulated to a toxic level in the shoots ahead of Na+. Our previous work suggested that although both Na+and Cl−are toxic to soybean under salt stress, there are differences between the sensitivity to Na+and Cl−forG. maxandG. soja(the wild soybean) [8,12,28]。Cl−有毒ity is more deleterious than Na+to the cultivated soybean and the damage is positively related to the Cl−contents within the leaves and stems, whereas the wild soybean and its hybrids with the cultivated soybean have stronger Cl−tolerance than the cultivated soybean itself [8,12,28]。
GmCLC1(GenBank accession: AY972079; Phytozome database: Glyma.05G077100) encodes a tonoplast-localized and pH-dependent Cl−/H+antiporter, which was upregulated by NaCl or dehydration stress, andGmCLC1-transgenic tobacco BY-2 cells displayed enhanced NaCl tolerance through increased Cl−transport from the cytoplasm into the vacuole [29,30]。Our prior study also found thatGmCLC1-transgenicArabidopsis thalianaandGmCLC1-overexpressing soybean hairy roots had enhanced Cl−/salt tolerance by reducing the Cl−accumulation in shoots or by sequestering more Cl−in roots [21]。The survival rate ofGmCLC1-transgenicΔgef1mutant yeast cells was also increased under varied levels of Cl−/salt stress [21]。Based on publishedGmCLC1information and public databases, we found seven other homologousCLCmembers,b1,b2, c1, c2, d1, d2 and g, in the soybean genome in both wild and cultivated soybean [22]。
In this work, all the homologous soybeanCLCmembers were cloned from the Cl−-sensitiveG. maxcultivar N23674 and the Cl−-tolerantG. sojaaccession BB52. By examining their expressions under favorable or NaCl-treated conditions, we isolatedGsCLC-c2, which is the only CLC member having non-synonymous changes betweenG. maxN23674 andG. sojaBB52. We have also systemically investigated its functions using hairy root-composite soybean plants, site-directed mutagenesis (SDM)-mediated yeast mutant complementation, and electrophysiological assays withXenopus laevisoocytes. Our goal is to decipher the molecular and physiological roles ofGsCLC-c2in soybean Cl−/salt stress adaptations, so as to provide an important theoretic basis for improving Cl−/salt tolerance in soybean and other crops.
Results
Whole-genome analyses ofCLChomologous genes inG. maxandG. soja
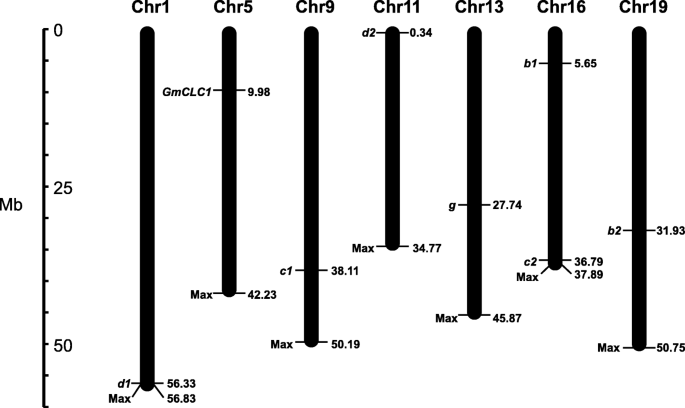
Using the Phytozome database (https://phytozome.jgi.doe.gov/pz/portal.html), we have previously found seven homologs ofGmCLC(Glyma.16G057600, Glyma.19G089800, Glyma.09G157900, Glyma.16G208400, Glyma.01G239000, Glyma.11G004600 and Glyma.13G161800) in the soybean genomes of both wild and cultivated soybeans, and named themb1,b2,c1,c2,d1,d2andg[22]。Here we extracted total RNA from the roots, stems and leaves ofG. maxcultivar N23674 andG. sojaaccession BB52 15-day-old seedlings, produced cDNAs by reverse transcription of mRNAs, and successfully cloned these seven newCLC同源染色体。These eight soybeanCLChomologs (includingGmCLC1) are located on chromosomes 1, 5, 9, 11, 13, 16 and 19, with two of the members,b1andc2, both being on chromosome 16 (Fig.1). The coding sequence (CDS) lengths of theseCLCsare between 2178 and 2472 bases. With the exceptions ofGmCLC1,c1, andd1(which are identical between the wild and cultivated soybeans), a few synonymous nucleotide substitutions were found in the otherCLCmembers betweenG. maxcultivar N23674 andG. sojaaccession. The only exception isc2where there is a 462 T > A substitution from cultivar N23674 to accession BB52, which resulted in the corresponding D154E change (Additional file1: Table S1).
Differential expressions ofCLCs inG. maxandG. sojaunder NaCl stress
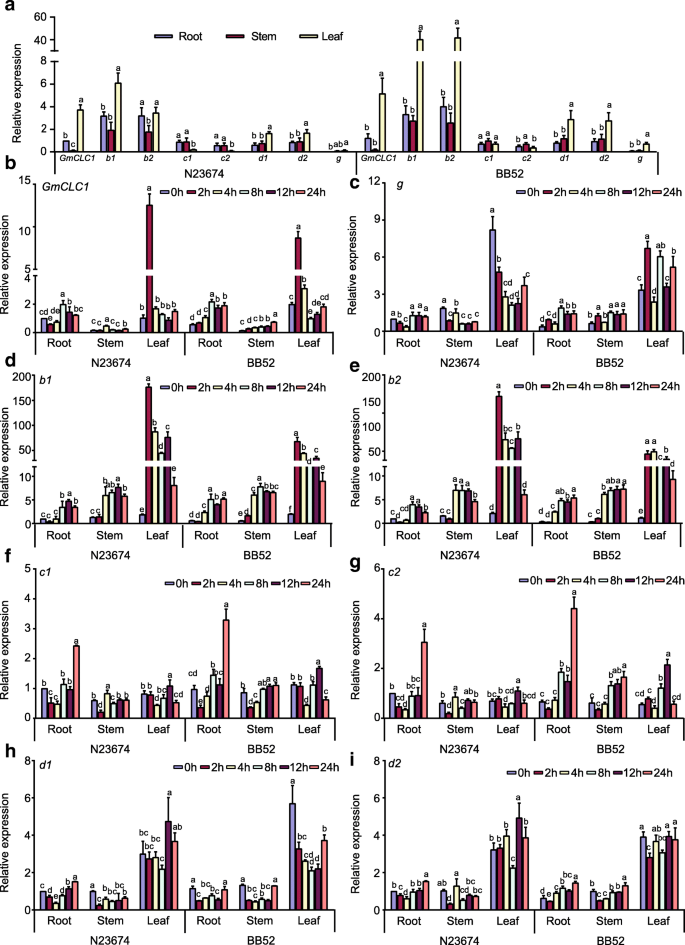
Under normal growth conditions, all eightCLChomologs were expressed in roots, stems and leaves of 15-day-old seedlings ofG. maxcultivar N23674 andG. sojaaccession BB52. Generally, the relative expressions in the leaves of most of theCLChomologs, includingGmCLC1[21,29,30],b1,b2,d1,d2andg, were markedly higher than those in the roots and stems of both wild and cultivated soybeans. However, the relative expressions ofc1andc2showed a different trend. Expression of bothc1andc2was lower in leaf than in root and stem of N23674 (Fig.2a). Expression ofc1in BB52 showed no significant difference in leaf, stem and root. But expression ofc2was significantly higher in stem compared to that in root and leaf (Fig.2a). When the plants were treated with 150 mM NaCl,GmCLC1,b1andb2were initially up-regulated (for the first 2 h) and were then down-regulated quickly in the leaves of both N23674 and BB52, whereas the expression of membergin the leaves declined gradually with increasing salt treatment time in N23674 but fluctuated in the leaves of BB52 (Fig.2b-e). The expressions ofGmCLC1,b1andb2were slightly up-regulated in the roots and stems of both wild and cultivated soybean seedlings under salt stress after 8 h (Fig.2b-e). With respect tod1andd2, their expressions decreased after the first 2 h in the roots and stems and after 8 h in the leaves, and then increased again with longer salt treatment time to about the same level as before the salt treatment (Fig.2h-i). On the other hand, the expressions ofc1andc2in the leaves peaked after 12 h of NaCl exposure and then dropped back to the level before salt treatment (Fig.2f-g). More significantly, their expressions in the roots increased more than 2-fold after 24 h of salt treatment, in both N23674 and BB52 (Fig.2f-g). The distinctively different transcriptional responses ofc1andc2, in the different tissues under both normal and salt-stress conditions and in both wild and cultivated soybeans, from those of the other soybeanCLCmembers make them warrant further investigation. Between the two,c2is the only one with a difference in amino acid sequences betweenG. maxcultivar N23674 andG. sojaaccession BB52 (Additional file1: Table S1). Therefore, we decided to examine in greater detail what physiological (including electrophysiological) and molecular roles the version ofc2found inG. sojaBB52 (heretofore to be namedGsCLC-c2) might play in soybean Cl−/salt tolerance.
Relative expressions ofCLC-homologous genes inG. maxN23674 andG. sojaBB52 15-day-old seedlings.aUnder normal conditions. All expressions were normalized to the expression ofGmCLC1in root of N23674. Expression ofGmCLC1in root in N23674 was set as 1 for comparison.b-iExpression of differentCLC-homologs under 150 mM NaCl stress for 0, 2, 4, 8, 12, and 24 h.g, b1, b2, c1, c2, d1andd2areCLC同源染色体。转录水平have been normalized against those ofGmEF1α2a. Expression of the gene in root of N23674 was set as 1 for comparison. The results were presented as mean ± SD (3 ≤ n ≤ 5). Letters indicate groups with statistically significant differences (P ≤ 0.05) using Duncan’s test after one-way ANOVA
Certain site-directed mutagenesis (SDM) mutants ofGsCLC-c2failed to complement the chloride channel-deficient yeast mutantΔgef1under NaCl, KCl, or KNO3treatment
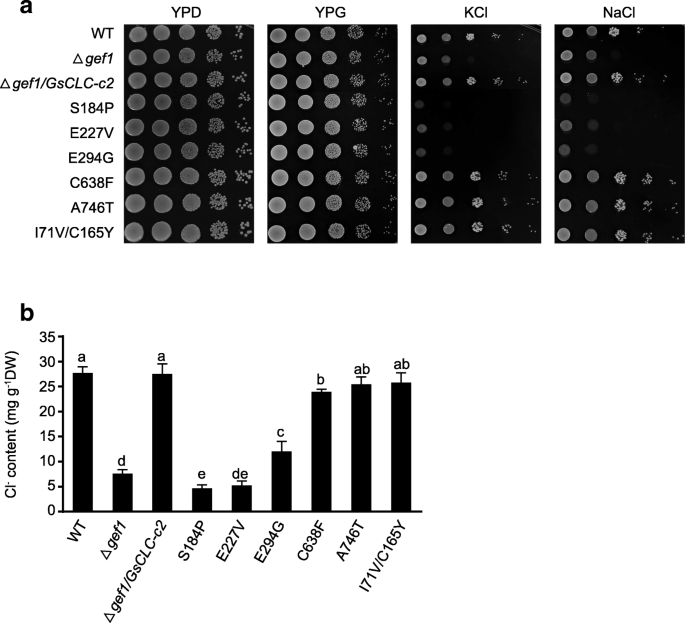
All the yeast strains, including the WT (BY4741),Δgef1(a GEF1-deficient, salt-sensitive mutant), andΔgef1transformed withGsCLC-c2or SDM mutants ofGsCLC-c2(S184P,E227V,E294G,C638F,A746TandI71V/C165Y), grew well on both YPD (yeast extract, peptone, dextrose) and YPG (yeast extract, peptone, galactose) media under favorable conditions. When cultured on YPG medium supplemented with 1 M KCl, or NaCl, theΔgef1mutant was unable to grow, but this phenotype was rescued by transformation with wild typeGsCLC-c2. However, mutants transformed withS184P,E227V, andE294G, with the key residues of the conserved domains I, II and III, respectively, ofGsCLC-c2mutated, failed to complement the growth defect of yeast cells in the presence of high concentrations of chloride salts. This suggested that these mutants could not mediate Cl−homeostasis in yeast, and therefore these specific residues may be important for the anion transport activity of GsCLC-c2. The otherGsCLC-c2mutants,C638F,A746T, andI71V/C165Y, however, could complement theΔgef1mutation, similar to theΔgef1mutants transformed with the wild typeGsCLC-c2(Fig.3a). The Cl−contents of WT,Δgef1/GsCLC-c2andΔgef1/GsCLC-c2mutants (C638F,A746T, andI71V/C165Y) grown in 1 M NaCl were significantly higher than those of theΔgeflmutants orΔgefmutants transformed withS184P,E227V, andE294G(P < 0.05), and theΔgefmutants transformed withS184PorE227Vhad similar Cl−levels as the untransformedΔgef1mutants (Fig.3b). This indicates that GsCLC-c2 has a similar function to the yeast GEF1 in Cl−transport, and that the S184P and E227V substitutions completely, and the E294G substitution partially, negated the Cl−transport function of GsCLC-c2.
Effects of expressing wild type or mutantGsCLC-c2on salt-sensitive yeast mutantΔgef1under salt treatments.aSurvival in YPD and YPG media, and YPG media supplemented with 1 M KCl or 1 M NaCl.bCl−contents in yeast cells under 1 M NaCl treatment. S184P, E227V, E294G, C638F, A746T and I71V/C165Y are mutants with specific amino acid substituted. YPD:1% yeast extract/2% peptone/2% dextrose; YPG:1% yeast extract/2% peptone/2% galactose. Each bar represents mean ± SD of ion contents of three independent cultures of each strain. Letters indicate groups with statistically significant differences (P ≤ 0.05) using Duncan’s test after one-way ANOVA
GsCLC-c2alleviates salt injury on transformed soybean hairy root composite plants by regulating the contents of Na+, K+, Cl−, NO3−and ratios of Cl−/NO3−and Na+/K+
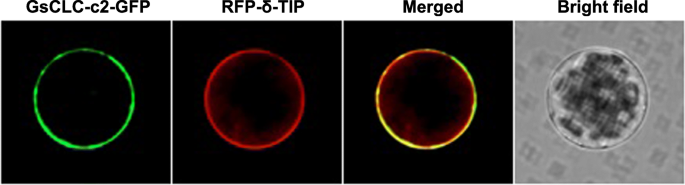
Using confocal microscopy, majority of GFP tagged GsCLC-c2 was found co-localized with the RFP tagged tonoplast marker protein δ-TIP (Fig.4) suggesting that GsCLC-c2 may serve similar functions as other tonoplast localized CLCs.
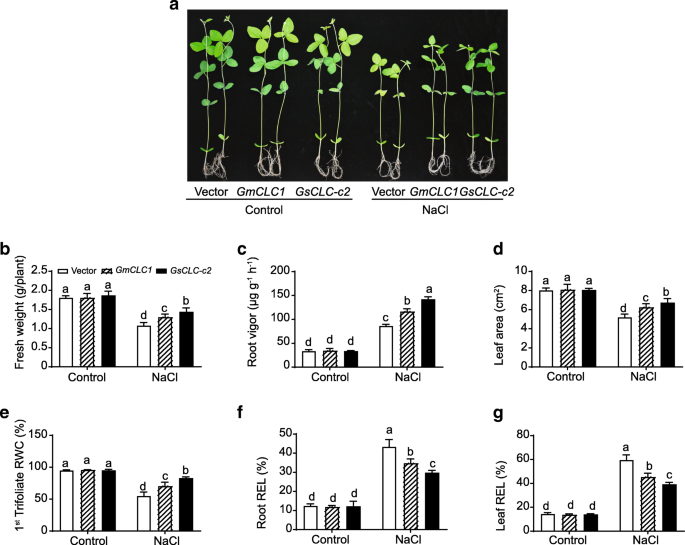
To confirm the function of GsCLC-c2, we adopted the soybean hairy root-composite plant system. Composite plants expressingGmCLC1were used for comparison. For each parameter, multiple comparisons were performed to compare the performance of all transgenic lines under control and treatment conditions using one-way ANOVA. Under favorable (control) conditions, the empty vector-transformed,GsCLC-c2-transformed andGmCLC1-transformed hairy root soybean (G. maxcv. N23674) composite plants all grew well with no observable phenotypic difference in term of fresh weight, root vigor, leaf area, relative water content (RWC) in first trifoliate, and relative electrolyte leakage (REL) in root and leaf (Fig.5). When subjected to 120 mM NaCl solution for 7 d, all parameters were significantly affected compared to the control condition regardless of the transgene. TheGsCLC-c2-transformed plants showed significantly better performance in fresh weight, root vigor, leaf area and RWC in the first trifoliate compared with the vector only control and theGmCLC1-transformed plants (Fig.5b-e). Plant transformed withGsCLC-c2also showed less REL suggesting that they were under less salt damage at cellular level in contrast to the vector only control and theGmCLC1-transformed plant (Fig.5f-g).
Comparisons of salt stress-related physiological parameters inGsCLC-c2- andGmCLC1-transgenic soybean hairy root-composite seedlings.aPhoto showing the hairy root-composite seedlings with or without NaCl treatment.bFresh weight, (c) root vigor, (d) leaf area, (e) relative water content (RWC) of the 1st trifoliate, (f) root relative electrolyte leakage (REL), and (g) leaf REL were compared between vector-transformed-,GsCLC-c2- andGmCLC1-transgenic soybean hairy root-composite seedling with or without NaCl treatment. Results are presented as mean ± SD (n = 3). Data of different lines under control and NaCl treatment were analyzed together by one-way ANOVA followed by Duncan’s test. Letters indicate groups with statistically significant differencesP ≤ 0.05
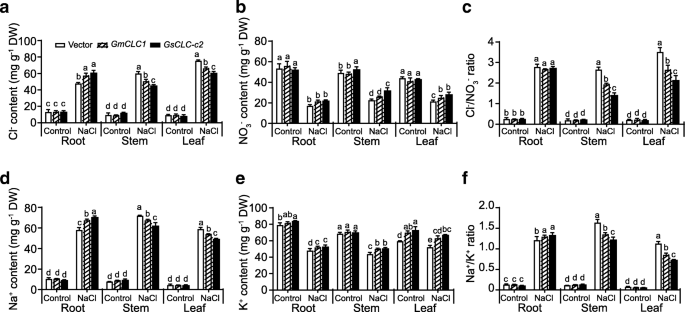
To confirm the salt tolerance was brought about by the primary functions ofGsCLC-c2, we also investigated the ion content in the transgenic composite plants. For each parameter, multiple comparisons were made to compare the performance of all transgenic lines in each tissue under control and treatment conditions using one-way ANOVA. Under control conditions, there was no significant difference in Cl−, Na+, Cl−/NO3−and Na+/K+ratio between the three constructs. Under the same conditions, composite plant ofGsCLC-c2showed significantly higher NO3−content in stem and K+content in root and leaf. Under salt stress, all lines showed elevation of Cl−and Na+contents but reduction in NO3−and K+contents. BothCLCs expressing plants showed higher Cl−content (accompanying with a higher Na+content) in the root but a reduction of Cl−content (accompanying with a reduction of Na+content) in the aerial parts (Fig.6a and d). With respect to NO3−, its level is higher in stem and leaf of composite plant ectopically expressingGsCLC-c2under salt stress conditions (Fig.6b). Accordingly, ectopic expression ofGsCLC-c2maintained significantly lower Cl−/NO3−and Na+/K+ratios in stem and leaf (Fig.6c-f). Again, like the physiological parameters,GsCLC-c2transgenic plants accumulated a significantly lower Cl−content in the aerial parts compared with the vector-transformed andGmCLC1-transformed plant (Fig.6a) suggested that GsCLC-c2, under salt stress, has stronger ion transportation activities over GmCLC1.
Ion content in different tissues of transgenic hairy root-composite plant.aCl−content, (b) NO3−content, (c) Cl−/NO3−ratio, (d) Na+content, (e) K+content, and (f) Na+/K+ratio of transgenic hairy root-composite plants. Composite plants with hairy root transformed with vector,GmCLC1, andGsCLC-c2were employed in this study. Results are presented as mean ± SD (n = 3). Data of different lines within the same tissue under control and NaCl treatment were analyzed together by one-way ANOVA followed by Duncan’s test. Letters indicate groups with statistically significant differencesP ≤ 0.05
Electrophysiological analyses ofXenopus laevisoocytes expressing GsCLC-c2
In order to further verify the anion transport activities of GsCLC-c2 to see if it is a bona fide anion channel protein, two-electrode voltage clamp experiments were performed usingXenopus laevisoocytes injected withGsCLC-c2cRNAs, and the electrophysiological track records for membrane steady-state current changes were conducted in vivo at 20 mV intervals between − 100 mV and + 100 mV. Oocytes microinjected with water (H2O) were used as the negative control to monitor the basal current. The results showed that, with the increase in voltage from − 100 mV to + 100 mV, the current increased at a much higher rate in theGsCLC-c2-transformed oocytes than the water-injected controls, reflecting a significant increase in the rate of change in conductance across the cell membrane. The invert rectifying current signified the movement of anion into or cation out of the oocytes. When the anion channel inhibitor 5-nitro-2-(3-phenylpropylamino) benzoic acid (NPPB) was added to the extracellular bath solution, the voltage-dependent change in current was not significantly different from the negative controls (H2O and H2O + NPPB) (Fig.7a). Therefore, this demonstrates that GsCLC-c2 possesses anion transport activity, and that anion is very likely Cl−, since NPPB is a well-known chloride channel blocker [31]。
Voltage clamp analyses ofGsCLC-c2expressions inXenopus laevisoocytes.aEffects of NPPB. Water- orGsCLC2-c2-cRNA injected oocytes were incubated in standard bath solution (96 mM NaCl) with or without the anion channel blocker NPPB.bEffects of Cl−concentrations.GsCLC2-c2cRNA injected oocytes were incubated in bath solution containing 0, 3, 30 or 96 mM NaCl.cpH effect.GsCLC2-c2cRNA injected oocytes were incubated in standard bath solution containing 96 mM NaCl at pH 5.5, 7.5, or 8.5.dHalide ion selectivity. Cells were kept in a bath solution containing 96 mM NaCl, NaBr or NaI. Results are presented as mean ± SD (n ≥ 4)
To confirm the voltage-dependent currents induced by GsCLC-c2 was mainly due to its Cl−transport activity, the currents across the membranes ofGsCLC-c2-transformed oocytes, kept in Cl−solutions at different concentrations (0, 3, 30, 96 mM), were recorded. The rate of increase in membrane currents was in direct proportion to the increase in ambient Cl−concentration (Fig.7b, Additional file1: Figure S1), indicating that the anion transport activity of GsCLC-c2 was positively correlated with Cl−concentrations. However, this activity was independent of pH at the levels tested (pH 5.5, 7.5 and 8.5) (Fig.7c). This indicates that the Cl−transport activity of GsCLC-c2 may not be coupled with H+exchange. In addition, to investigate the affinity of GsCLC-c2 for different halide ions,GsCLC-c2-transformed oocytes were subjected to the standard bath solutions containing Br−, I−or Cl−at the same concentration. A voltage-dependent conductance response curve was obtained for all three ions, and the conductance responses to Br−and I−were about the same, and both were greater than that to Cl−(Fig.7d).
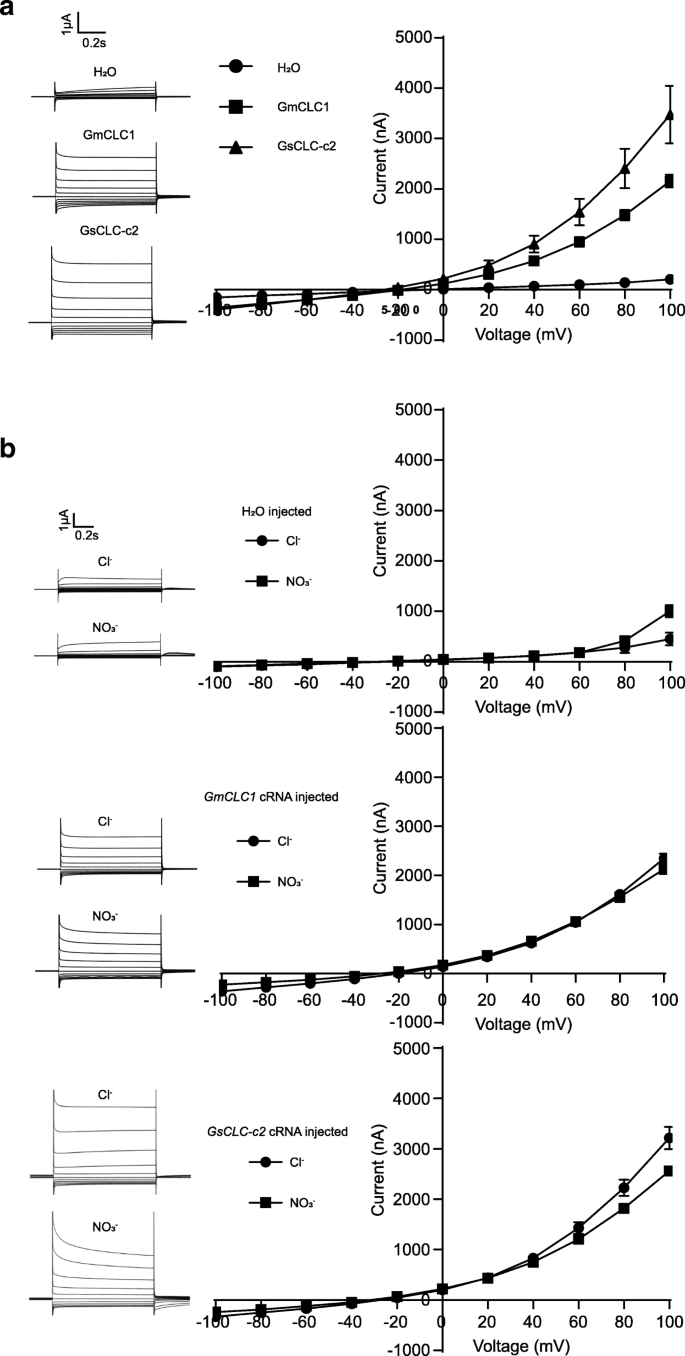
Because of the discrepancy between the anion transport activity of GsCLC-c2 reported here and that of the previously reported GmCLC1 with respect to pH dependence, we decided to directly compare the Cl−transport activity between GsCLC-c2 and GmCLC1. BothGsCLC-c2-transformed as well asGmCLC1-transformed oocytes responded to the extracellular Cl−bath solutions with significantly higher rates of change in conductance with increasing voltage compared to water control. However, the increase in currents induced by GsCLC-c2 was significantly steeper than that by GmCLC1, which demonstrates that the Cl−transport activity of GsCLC-c2 is higher than that of GmCLC1 (Fig.8a, Additional file1: Figure S2). WhenGsCLC-c2-cRNA-transformed as well asGmCLC1-cRNA-transformed oocytes were exposed to either Cl−or NO3−bath solutions with the same anion concentrations,GmCLC1-cRNA-transformed oocytes did not show any significant difference between the current response curves to Cl−and to NO3−, but the response curve for Cl−was significantly steeper than that for NO3−in theGsCLC-c2-transformed oocytes. The signal also attenuated faster for NO3−than for Cl−with GsCLC-c2 (Fig.8b). This indicates that, different from GmCLC1, GsCLC-c2 shows a slightly higher affinity for Cl−than for NO3−.
Electrophysiological comparisons ofXenopus laevisoocytes expressingGsCLC-c2andGmCLC1.aIon transport activity in a bath solution containing 96 mM NaCl.bCl−and NO3−selectivity in a bath solution containing 96 mM NaCl or and 96 mM NaNO3, respectively. Results are presented as mean ± SD (n ≥ 5). Upper panel: oocytes were injected with water; middle panel: oocytes were injected withGmCLC1cRNA; lower panel: oocytes were injected withGsCLC-c2cRNA
Discussion
Investigations into Cl−有毒ity have been relatively sparse compared to those on the effects of Na+on plants under salt stress [11,14,20,32]。One of the few examples is the discovery that the rootstocks from salt-tolerant species of citrus and grape played a critical role in enhancing the Cl−/salt tolerance of the “chloride-sensitive” or “chloride-hating” seedlings [33,34,35]。Other studies have found that the key steps for alleviating Cl−有毒ity involve the sequestering of Cl−in the root xylem parenchyma cells to limit its transport from the roots to the aerial parts, and the efflux of Cl−by the root stele cells or compartmentalization of Cl−in the vacuoles to reduce its availability in the roots [11,36]。CLCs can play important roles in these processes and contribute to Cl−homeostasis and Cl−/salt tolerance of plants [24,32,37,38]。In Arabidopsis, there are seven members in the CLC protein family but only AtCLCc and AtCLCg (with 62% amino acid sequence identity to each other) have been shown to participate in Cl−homeostasis, stomatal movement and Cl−/salt tolerance [24,32,37,39]。The ectopic expression ofCsCLCcfrom trifoliate orange (Citrus trifoliata) inAtclc-cmutant plants displayed improved seed germination, better growth and lower Cl−accumulation in the roots and shoots than the non-transgenic mutant seedlings under salt stress [38]。
In this study, the other seven members besidesGmCLC1(b1,b2, c1, c2, d1, d2andg) of the soybeanCLCgene family found in bothG. max(cultivar N23674) andG. soja(accession BB52) have been successfully cloned for the first time and their sequences verified (Fig.1, Additional file1: Table S1). Specifically,c2is the only one that has an amino acid change between the cultivated (cultivar N23674) and the wild soybean (accession BB52): D154E, through a single nucleotide substitution (c.462 T > A) (Additional file1: Table S1). At present, it is not clear whether this specific variation alone is responsible for the Cl−/salt tolerance differences between the cultivated and wild soybeans. However, based on the unique patterns of expressions ofc1andc2in the roots, stems and leaves of N23674 and BB52 seedlings under both favorable and salt-stress conditions (Fig.2), we postulate thatGsCLC-c2may be more responsible for soybean responses to Cl−/NaCl stress in root than otherCLCmembers. Similar to GmCLC1, GsCLC-c2 is also localized to the tonoplast (Fig.4). Under NaCl stress, the salt damage toGsCLC-c2-overexpressing毛茸茸的根composite soybean plants like theGmCLC1-overexpressing ones was clearly alleviated when compared to the empty vector-transformed plants (Fig.5a). If we just look at the ionic effects of salt injury, this alleviation was correlated with the significant reduction in Cl−contents of the stems and leaves of salt-stressed soybean plants, and indirectly related to the maintenance of NO3−accumulation in various parts of the plants [9,21,22]。These combined phenomena can effectively reduce the Cl−/NO3−ratios in the stems and leaves of the plants (Fig.6c). Based on the results from theΔgef1yeast mutant assays (Fig.3) and its localization on the tonoplast ofArabidopsisprotoplasts (Fig.4), we postulate that GsCLC-c2 probably affects this by increasing the sequestration of Cl−into the root cell vacuoles. Similarly, under salt stress, K+contents in the stems and leaves were increased, and Na+levels and Na+/K+ratios were both reduced in the aerial parts while raised in the roots ofGsCLC-c2-overexpressing毛茸茸的根composite soybean plants, implying more Na+has been sequestered in the roots of these transgenic plants than in the controls (Fig.6d-f). It is largely unknown how CLC regulates the Na+/K+ratio. Yet, it has been suggested that Cl−and Na+transportation is somehow coupled [14]。Alteration of one would sometimes result in the alteration of the other [27,40]。Furthermore, whetherGsCLC-c2is directly involved in the Cl−accumulation or sequestration in the vacuoles of the roots of the transgenic composite soybean plants under salt stress, or whether it indirectly contributes to the restriction of Cl−transport from roots to shoots requires further investigations.
In Arabidopsis, whether the anion channel proteins, AtCLCa-d and AtCLCg, have a higher affinity for NO3−or Cl−is determined by the amino acid “x” in the highly conserved domain I – GxGIPE. If x is P (proline), NO3−would be the preferred anion; if x is S (serine), Cl−would be preferred [24,26]。GmCLC1, with 78% identity to AtClCa, is the first CLC reported in soybean and has a proline residue (P165) in the conserved domain I [23,26]。Our previous study showed that GsCLC-c2 has a serine residue (S184) in the same conserved domain [23], and here we demonstrated that GmCLC1 did not show a preference between Cl−and NO3−whereas GsCLC-c2 had a slightly higher response to Cl−over NO3−(Fig.8b).
Heterologous expression in special yeast mutants has been exploited by many researchers aiming to functionally characterize ion transporters from plant [21,29,30,41,42]。The GEF1-deficient yeast mutant (Δgef1)-based system was used to determine the efficacy ofGmCLC1in its ability to alleviate chloride/salt stress or its protective functions under Cl−stress [21]。In this study, the soybeanGsCLC-c2gene was able to complement the salt-sensitive phenotype of the△gef1mutant grown in YPG media containing KCl or NaCl (Fig.3a). When GsCLC-c2 was mutated by one amino acid substitution (S184P, E227V, and E294G in the conserved domains I, II, and III, respectively), its ability to complement thegef1mutation in yeast was very much abolished. The ability of the S184P-transformed yeast mutant to accumulate Cl−in the cells was also drastically reduced (Fig.3b), which could be the result of changing the Cl−or NO3−specificity by the presence of either serine or proline in the determining position within the conserved domain I. However, the other mutations, such as C638F, A746T and I71V/C165Y within the non-conserved areas had no effect on the ability of GsCLC-c2 to complement the restrained growth of the△gef1mutant in media containing KCl, NaCl (Fig.3). The△gef1mutant transformed with these mutated forms ofGsCLC-c2also accumulated comparable levels of Cl−to those transformed with the wild typeGsCLC-c2(Fig.3b), indicating that these non-conserved domains may not play any part in the protein’s function as an anion/chloride channel. This result also implied that introducing these amino acid substitutions has little effect on the function of this protein by affecting its conformation (Fig.3).
TheX. laevisoocyte, a heterologous expression system, has been successfully used to study various biological processes and ion channels [30,43]。In this work, GsCLC-c2-expressingX. laevisoocytes displayed a significantly elevated voltage-dependent conductance when bathed in a solution containing 96 mM NaCl, and this was negated by the inhibitory effect of the chloride channel blocker, NPPB (Fig.7a). Furthermore, the conductance exhibited by the GsCLC-c2-expressing oocytes was elevated in proportion to increasing Cl−concentrations (Fig.7b, Additional file1: Figure S1), but, surprisingly, independent of pH (Fig.7c). The latter indicates that the Cl−transport function of GsCLC-c2 does not involve H+exchange, and this is not consistent with the functions of AtCLCa,which can transport Cl−or NO3−reversely coupled with H+transport, and its affinity for NO3−is stronger than for Cl−[43,44]。GsCLC-c2 is also different from GmCLC1, whose anion transport function showed pH dependence[30]。It appears that GsCLC-c2 is an anion channel and not a H+/anion cotransporter as deduced from our previous bioinformatics analyses [23]。Therefore, further studies of the functions of GsCLC-c2 are required.
Conclusions
GsCLC-c2is up-regulated in the roots of NaCl-stressed wild soybean plants, and the encoded protein is more effective than GmCLC1 as a chloride channel, with higher permeability to Cl−and higher affinity for Cl−than NO3−than GmCLC1, and is responsive to halide ions in general. Its chloride transport function is pH-independent, but dependent on the amino acid sequences in the conserved domains I, II and III. It is localized in the tonoplast and therefore likely contributes to enhanced Cl−/salt tolerance in plants by increasing the sequestration of excess Cl−in the vacuoles of root cells and thus preventing Cl−from being transported to the shoots where it can result in cellular damages.
Methods
Plant materials, bacteria and yeast strains,Xenopus laevisoocytes, and plasmids
Plant seeds, includingGlycine max(L.) Merr. cultivar N23674 (salt-sensitive) were obtained from National Center for Soybean Improvement, Key Laboratory of Biology and Genetic Improvement of Soybean, Nanjing Agricultural University, Nanjing, China. Seeds ofG. sojaaccession BB52 (salt-tolerant) were collected from the coastal area in Shandong province of P.R. China [8,12]。No special permissions were required to use this wild soybean accession. Wild-type (WT)Arabidopsis thalianaColumbia-0glabrous1–1(Col-0gl1–1, alias Col-5) was a gift from Dr. H.Z. Shi lab of Texas Tech University, USA.Escherichia coliDH5α,Agrobacterium rhizogenesstrain K599, plant transient expression vectors pJIT166-GFP and pJIT166-RFP, binary vector for plant transformation pCAMBIA1300, yeast mutantΔgef1in BY4741 background and yeast expression plasmid pYES2,Xenopus laevis卵母细胞和质粒pGEHME代cRNA were used in this study.
Cloning of soybeanCLCgenes and vector construction for plant, bacterial and yeast transformation
The seed germination and seedling cultivation of soybean were conducted as previously reported [21]。从15天大大豆提取的总RNAseedlings using the TaKaRa MiniBEST Plant RNA Extraction Kit (TaKaRa, Dalian, China). The full-length coding sequence (CDS) ofCLCgenes were amplified in PCR reactions containing 1× PCR buffer, 0.15 mM MgCl2, 0.25 mM dNTPs, 0.2 μM of each primer, 0.25 U KOD-Plus DNA polymerase (TOYOBO, Japan), and 2 μL first-strand cDNA produced with PrimeScript™ II 1st Strand cDNA Synthesis Kit (TaKaRa, Dalian, China) according to the manufacturer’s protocol. The PCR products were ligated to the pMD 19-T vector (TaKaRa, Dalian, China) and sequencing of the clones was done using the sequencing primer sites on the vector. Primer sequences are listed in Additional file1: Table S2.
The CDS ofGsCLC-c2without the stop codon was cloned upstream of and in-frame with the CDS of green fluorescent protein (GFP) in the pJIT166-GFP plasmid to obtain the recombinant plasmid pJIT166-GsCLC-c2-GFP. The tonoplast marker gene,δ-TIP, for co-localization was cloned in-frame with the 3′-terminus of red fluorescent protein (RFP) CDS in the pJIT166-RFP plasmid to obtain pJIT166-RFP-δ-TIPfor Arabidopsis leaf protoplast transformation [39,45]。The CDS ofGsCLC-c2was cloned into pCAMBIA1300 to obtain the pCAMBIA1300-GsCLC-c2, which was then transformed intoA. rhizogenesK599 for plant transformation. To expressGsCLC-c2in the yeast mutantΔgef1, the CDS was cloned into the yeast expression vector pYES2. To expressGsCLC-c2andGmCLC1inX. laevisoocytes, their CDS were cloned into pGEHME. Primers used for amplifying the above genes are shown in Additional file1: Table S2. pCAMBIA1300-GmCLC1was constructed as our previous work [21]。
Chromosomal location determination
In order to pinpoint the distribution ofGmCLCsfamily members throughout the soybean genome, we used each member as a query against the soybean genome on theSoyBase Wm82 Genome Browser(https://www.soybase.org/gb2/gbrowse/gmax2.0/) to determine the location of the transcription initiation site of each gene. The MapInspect software (version 1.0) was used to draw the locations ofGmCLCgenes on each chromosome.
qRT-PCR assays
The roots, stems, and leaves were sampled from 15-day-old N23674 and BB52 plants treated with 150 mM NaCl for 0, 2, 4, 8, 12, and 24 h, respectively, and were frozen in liquid N2RNA的提取。总rna是孤立的使用MiniBEST Plant RNA Extraction Kit (TaKaRa, Dalian, China) and then used for the 1st strand cDNA synthesis following the instructions of PrimeScript RT reagent Kit with gDNA Eraser (TaKaRa). A soybean elongation factor 1-alpha-like gene,EF1α2a(XM_003524541.1) [46], was used as an internal reference. qRT-PCR reactions were performed in 96-well plates using the StepOnePlus Real-Time PCR System (ThermoFisher Scientific China, Inc., Shanghai) with SYBR® Premix Ex Taq™ II (TaKaRa) according to manufacturer’s protocols in a 10-μL reaction. Data were analyzed by StepOnePlus Software v2.1 (Thermo Fisher Scientific China, Inc., Shanghai). The relative expression of each gene was normalized against the internal reference gene, and calculated according to the 2−ΔΔCTmethod [47]。Primers used in the qRT-PCR are presented in Additional file1: Table S2.
Subcellular co-localization assays of GsCLC-c2
The seed germination and seedling cultivation of Arabidopsis were conducted as previously reported [21]。Arabidopsis protoplasts were isolated from the leaves of 25-day-old plants as previously reported [48]。Two fusion constructsGsCLC-c2-GFPand tonoplast specific markerRFP-δ-TIP[39] were co-transformed into the protoplast by the PEG4000-mediated method [49]。After incubation of the transformed Arabidopsis protoplasts for 18–24 h at room temperature, GFP and RFP signals were detected by confocal fluorescence microscopy (PerkinElmer UltraVIEW VoX).
Site-directed mutagenesis (SDM)-mediated complementation tests ofGsCLC-c2in yeast mutant
To investigate the Cl−transport functions of GsCLC-c2, site-directed mutagenesis ofGsCLC-c2was carried out. The strategies for the SDM ofGsCLC-c2are shown in Additional file1: Table S3, and the mutations were performed with the Muta-direct™ site-directed mutagenesis Kit (SBS Genetech, Shanghai, China) using the pYES2-GsCLC-c2construct according to manufacturer’s protocol. The desired constructs confirmed with sequencing were transformed into the salt-sensitive yeast mutant△gef1as described previously [50]。The yeast cultures in 10-fold dilution series beginning at OD600 ≈ 0.5 were plated on YPD medium (1% yeast extract/2% peptone/2% dextrose), YPG medium (1% yeast extract/2% peptone/2% galactose), with or without 1 M NaCl, or 1 M KCl supplement. The plates were photographed 2~4 days after incubation at 30 °C. Cl−contents of WT (S. cerevisiaeBY4741),Δgef1mutant,Δgef1/GsCLC-c2, orΔgef1/GsCLC-c2mutants were determined with cultures grown in liquid YPG medium with 1 M NaCl. Cells were harvested at OD600 ≈ 0.2 by filtration, and their Cl−content was determined [51]。
Salt tolerance assays ofGsCLC-c2overexpressing hairy root composite soybean plants
Hairy root transformation was performed as described in our previous work [21]。Soybean seedlings were transformed with pCAMBIA1300 (Vector), pCAMBIA1300-GsCLC-c2or pCAMBIA1300-GmCLC1usingA. rhizogenesstrain K599. Ten-day after K599 infection, seedlings with similar length of hairy roots were transferred to ½ X Hoagland solution with or without 120 mM NaCl. Solution was replaced every 3 days. Roots, stems and leaves were harvested 7-day after transfer for Na+, K+, Cl−and NO3−content measurement [51,52]。Photos of plants were taken right before sample harvest.
Xenopus laevisoocyte transformation withGsCLC-c2by microinjection and electrophysiology
Ambion mMESSAGE mMACHINE™ T7 Transcription Kit (AM1344, Life Technologies, California, United States) was used to synthesize capped cRNAs from linearized pGEHME-GsCLC-c2and pGEHME-GmCLC1. Harvesting and handling ofX. laevisoocytes were done as previously described [30]。Oocytes were injected with either 50 nL of H2O or 50 nL of 1 ng L− 1cRNA. The injected oocytes were incubated in the standard bath solution (96 mM NaCl, 2 mM potassium gluconate, 5 mM calcium D-gluconate, 1.2 mM MgSO4, 5 mM HEPES, pH 7.5) at 16~18 °C [43] before two-electrode voltage clamping. Data acquisition and analysis were done using the oocyte clamp recording interface (Warner Instrument OC-725C, Hamden, CT, USA) and pClamp9 software (Molecular Devices, San Jose, CA, USA) [30]。To verify whether GsCLC-c2 is a bona fide Cl−transporter, oocytes expressing the transporter were challenged with bath solution with different concentration of Cl−and 10 μM NPPB (5-nitro-2-[3-phenylpropylamino] benzoic acid, an anion channel inhibitor) [31]。In order to determine the effect of extracellular pH on the transporter function in the range of 5.5 to 8.5, the bath solution was buffered with 5 mM MES for pH 5.5, and with 5 mM Tris for pH 8.5. In ion substitution experiments, 96 mM NaCl was substituted with equal concentrations of NaI, NaBr or NaNO3. Ag/AgCl electrodes and 3 M KCl agar bridges were used as reference and bath electrodes, respectively [30]。
Statistical analyses
All data were analyzed and presented as the means ± SD for each treatment (n = 3; except for electrophysiology wheren = 4~9) using SPSS software (ver. 20.0). The data were subjected to the one-way analysis of variance (ANOVA), and pairwise comparisons were performed using Duncan’s test atP ≤ 0.05. For physiological data, data of different transgenic lines under control and NaCl treatment conditions were compared. For ion content, data of different transgenic lines among the same tissue under control and NaCl treatment conditions were compared.
Abbreviations
- ANOVA:
-
Analysis of variance
- CDS:
-
Coding sequence
- CLC:
-
Chloride channel
- Col-0gl1–1:
-
Arabidopsis thalianaColumbia-0glabrous1–1
- GFP:
-
Green fluorescent protein
- NPPB:
-
5-nitro-2-(3-phenylpropylamino) benzoic acid
- PSII:
-
Photosystem II
- REL:
-
Relative electrolyte leakage
- RFP:
-
Red fluorescent protein
- RWC:
-
Relative water content
- SDM:
-
Site-directed mutagenesis
- WT:
-
Wild-type
- YPD:
-
Yeast extract, peptone, dextrose
- YPG:
-
Yeast extract, peptone galactose
References
- 1.
Deinlein U, Stephan AB, Horie T, Luo W, Xu G, Schroeder JI. Plant salt-tolerance mechanisms. Trends Plant Sci. 2014;19(6):371–9.
- 2.
Ondrasek G, Rengel Z, Veres S: Soil salinisation and salt stress in crop production. In: Abiotic stress in plants - mechanisms and adaptations.Edited by Shanker A. Rijeka, Croatia InTech; 2011: 171–190.
- 3.
Roy SJ, Negrao S, Tester M. Salt resistant crop plants. Curr Opin Biotech. 2014;26:115–24.
- 4.
Genc Y, Oldach K, Taylor J, Lyons GH. Uncoupling of sodium and chloride to assist breeding for salinity tolerance in crops. New Phytol. 2016;210(1):145–56.
- 5.
Ismail AM, Horie T. Genomics, physiology, and molecular breeding approaches for improving salt tolerance. Annu Rev Plant Biol. 2017;68:405–34.
- 6.
Munns R, Tester M. Mechanisms of salinity tolerance. Annu Rev Plant Biol. 2008;59:651–81.
- 7.
Hasegawa PM. Sodium (Na+) homeostasis and salt tolerance of plants. Environ Exp Bot. 2013;92:19–31.
- 8.
Zhang XK, Zhou QH, Cao JH, Yu BJ. Differential cl−/salt tolerance and NaCl-induced alternations of tissue and cellular ion fluxes inGlycine max,Glycine sojaand their hybrid seedlings. J Agron Crop Sci. 2011;197(5):329–39.
- 9.
Guo JS, Zhou Q, Li XJ, Yu BJ, Luo QY. Enhancing NO3−supply confers NaCl tolerance by adjusting cl−uptake and transport inG. max&G. soja. J Soil Sci Plant Nut. 2017;17(1):194–204.
- 10.
Henderson SW, Baumann U, Blackmore DH, Walker AR, Walker RR, Gilliham M. Shoot chloride exclusion and salt tolerance in grapevine is associated with differential ion transporter expression in roots. BMC Plant Biol. 2014;14:273.
- 11.
Li B, Qiu J, Jayakannan M, Xu B, Li Y, Mayo GM, Tester M, Gilliham M, Roy SJ. AtNPF2.5 modulates chloride (cl−) efflux from roots ofArabidopsis thaliana. Front Plant Sci. 2017;7.https://doi.org/10.3389/fpls.2016.02013.
- 12.
Luo QY, Yu BJ, Liu YL. Differential sensitivity to chloride and sodium ions in seedlings ofGlycine maxandG. sojaunder NaCl stress. J Plant Physiol. 2005;162(9):1003–12.
- 13.
Qu YN, Zhou Q, Yu BJ. Effects of Zn2+and niflumic acid on photosynthesis inGlycine sojaandGlycine maxseedlings under NaCl stress. Environ Exp Bot. 2009;65(2–3):304–9.
- 14.
Teakle NL, Tyerman SD. Mechanisms of cl−transport contributing to salt tolerance. Plant Cell Environ. 2010;33(4):566–89.
- 15.
Wege S, Gilliham M, Henderson SW. Chloride: not simply a ‘cheap osmoticum’, but a beneficial plant macronutrient. J Exp Bot. 2017;68(12):3057–69.
- 16.
Franco-Navarro JD, Brumos J, Rosales MA, Cubero-Font P, Talon M, Colmenero-Flores JM. Chloride regulates leaf cell size and water relations in tobacco plants. J Exp Bot. 2016;67(3):873–91.
- 17.
Chen CZ, Lv XF, Li JY, Yi HY, Gong JM. Arabidopsis NRT1.5 is another essential component in the regulation of nitrate reallocation and stress tolerance. Plant Physiol. 2012;159(4):1582–90.
- 18.
O'Brien JA, Vega A, Bouguyon E, Krouk G, Gojon A, Coruzzi G, Gutierrez RA. Nitrate transport, sensing, and responses in plants. Mol Plant. 2016;9(6):837–56.
- 19.
Cubero-Font P, Maierhofer T, Jaslan J, Rosales MA, Espartero J, Diaz-Rueda P, Muller HM, Hurter AL, AL-Rasheid KAS, Marten I, et al. Silent S-type anion channel subunit SLAH1 gates SLAH3 open for chloride root-to-shoot translocation. Curr Biol. 2016;26(16):2213–20.
- 20.
Qiu JE, Henderson SW, Tester M, Roy SJ, Gilliham M. SLAH1, a homologue of the slow type anion channel SLAC1, modulates shoot cl−accumulation and salt tolerance inArabidopsis thaliana. J Exp Bot. 2016;67(15):4495–505.
- 21.
Wei PP, Wang LC, Liu AL, Yu BJ, Lam HM. GmCLC1 confers enhanced salt tolerance through regulating chloride accumulation in soybean. Front Plant Sci. 2016;7.https://doi.org/10.3389/fpls.2016.01082.
- 22.
刘于BJ, Z,魏页。在整个研究进展genome discovery and function analysis ofCLChomologous genes in plants based on salt stress. Journal of Nanjing Agricultural University. 2017;40(2):187–94.
- 23.
Li W, Wang LC, Cao JH, Yu BJ. Bioinformatics analysis of CLC homologous genes family in soybean genome. Journal of Nanjing Agricultural University. 2014;37(3):35–43.
- 24.
Barbier-Brygoo H, De Angeli A, Filleur S, Frachisse JM, Gambale F, Thomine S, Wege S. Anion channels/transporters in plants: from molecular bases to regulatory networks. Annu Rev Plant Biol. 2011;62:25–51.
- 25.
Wege年代,Jossier M, Filleur年代,Thomine年代,Barbier-Brygoo H, Gambale F, De Angeli A. The proline 160 in the selectivity filter of theArabidopsisNO3−/H+exchanger AtCLCa is essential for nitrate accumulationin planta. Plant J. 2010;63(5):861–9.
- 26.
Zifarelli G, Pusch m . CLC运输蛋白质在plants. FEBS Lett. 2010;584(10):2122–7.
- 27.
刘Lam HM,徐X, X,陈WB,杨GH,黄FL,李MW, He WM, Qin N, Wang B, et al. Resequencing of 31 wild and cultivated soybean genomes identifies patterns of genetic diversity and selection. Nat Genet. 2010;42(12):1053–U1041.
- 28.
Chen XQ, Yu BJ. Ionic effects of Na+and cl−on photosynthesis inGlycine maxseedlings under isoosmotic salt stress. Zhi Wu Sheng Li Yu Fen Zi Sheng Wu Xue Xue Bao. 2007;33(4):294–300.
- 29.
Li WYF, Wong FL, Tsai SN, Phang TH, Shao GH, Lam HM. Tonoplast-located GmCLC1 and GmNHX1 from soybean enhance NaCl tolerance in transgenic bright yellow (BY)-2 cells. Plant Cell Environ. 2006;29(6):1122–37.
- 30.
Wong TH, Li MW, Yao XQ, Lam HM. The GmCLC1 protein from soybean functions as a chloride ion transporter. J Plant Physiol. 2013;170(1):101–4.
- 31.
Barbier-Brygoo H, Vinauger M, Colcombet J, Ephritikhine G, Frachisse J, Maurel C. Anion channels in higher plants: functional characterization, molecular structure and physiological role. Biochim Biophys Acta. 2000;1465(1–2):199–218.
- 32.
Nguyen CT, Agorio A, Jossier M, Depre S, Thomine S, Filleur S. Characterization of the chloride channel-like, AtCLCg, involved in chloride tolerance inArabidopsis thaliana. Plant Cell Physiol. 2016;57(4):764–75.
- 33.
Alvarez-Gerding X, Cortes-Bullemore R, Medina C, Romero-Romero JL, Inostroza-Blancheteau C, Aquea F, Arce-Johnson P. Improved salinity tolerance in Carrizo citrange rootstock through overexpression of glyoxalase system genes. Biomed Res Int. 2015.
- 34.
Brumos J, Colmenero-Flores JM, Conesa IzquierdoP, Sanchez G, Iglesias DJ, Lopez-Climent MF, Gomez-Cadenas A, Talon M. Membrane transporters and carbon metabolism implicated in chloride homeostasis differentiate salt stress responses in tolerant and sensitiveCitrusrootstocks. Funct Integr Genomic. 2009;9(3):293–309.
- 35.
Tregeagle JM, Tisdall JM, Tester M, Walker RR. Cl−uptake, transport and accumulation in grapevine rootstocks of differing capacity for cl−-exclusion. Funct Plant Biol. 2010;37(7):665–73.
- 36.
Abbaspour N, Kaiser B, Tyerman S. Chloride transport and compartmentation within main and lateral roots of two grapevine rootstocks differing in salt tolerance. Trees-Struct Funct. 2013;27(5):1317–25.
- 37.
Jossier M, Kroniewicz L, Dalmas F, Le Thiec D, Ephritikhine G, Thomine S, Barbier-Brygoo H, Vavasseur A, Filleur S, Leonhardt N. TheArabidopsisvacuolar anion transporter, AtCLCc, is involved in the regulation of stomatal movements and contributes to salt tolerance. Plant J. 2010;64(4):563–76.
- 38.
Wei QJ, Liu YZ, Zhou GF, Li QH, Yang CQ, Peng SA. Overexpression ofCsCLCc, a chloride channel gene fromPoncirus trifoliata, enhances salt tolerance inArabidopsis. Plant Mol Biol Rep. 2013;31(6):1548–57.
- 39.
Lv QD, Tang RJ, Liu H, Gao XS, Li YZ, Zheng HQ, Zhang HX. Cloning and molecular analyses of theArabidopsis thalianachloride channel gene family. Plant Sci. 2009;176(5):650–61.
- 40.
Liu Y, Yu LL, Qu Y, Chen JL, Liu XX, Hong HL, Liu ZX, Chang RZ, Gilliham M, Qiu LJ, et al. GmSALT3, which confers improved soybean salt tolerance in the field, increases leaf cl- exclusion prior to Na+ exclusion but does not improve early vigor under salinity. Front Plant Sci. 2016;7.
- 41.
Shi H, Lee B-h, Wu S-J, Zhu J-K. Overexpression of a plasma membrane Na+/H+antiporter gene improves salt tolerance inArabidopsis thaliana. Nat Biotechnol. 2002;21:81.
- 42.
Zhao XF, Wei PP, Liu Z, Yu BJ, Shi HZ. Soybean Na+/H+antiporter GmsSOS1 enhances antioxidant enzyme activity and reduces Na+accumulation inArabidopsisand yeast cells under salt stress. Acta Physiol Plant. 2017;39(1).
- 43.
Bergsdorf EY, Zdebik AA, Jentsch TJ. Residues important for nitrate/proton coupling in plant and mammalian CLC transporters. J Biol Chem. 2009;284(17):11184–93.
- 44.
De Angeli A, Monachello D, Ephritikhine G, Frachisse JM, Thomine S, Gambale F, Barbier-Brygoo H. The nitrate/proton antiporter AtCLCa mediates nitrate accumulation in plant vacuoles. Nature. 2006;442(7105):939–42.
- 45.
Dong CH, Rivarola M, Resnick JS, Maggin BD, Chang C. Subcellular co-localization ofArabidopsisRTE1 and ETR1 supports a regulatory role for RTE1 in ETR1 ethylene signaling. Plant J. 2008;53(2):275–86.
- 46.
Saraiva KDC, de Melo DF, Morais VD, Vasconcelos IM, Costa JH. Selection of suitable soybeanEF1αgenes as internal controls for real-time PCR analyses of tissues during plant development and under stress conditions. Plant Cell Rep. 2014;33(9):1453–65.
- 47.
Schmittgen TD, Livak KJ. Analyzing real-time PCR data by the comparative CT method. Nat Protoc. 2008;3:1101.
- 48.
Schapire AL, Lois LM: A simplified and rapid method for the isolation and transfection ofArabidopsisleaf mesophyll protoplasts for large-scale applications. In: Plant signal transduction: methods and protocols. Edited by Botella JR, Botella MA, vol. 1363, 2nd edn. New York, NY: Humana Press; 2016: 79–88.
- 49.
Scala V, Grottoli A, Cigliano RA, Anzar I, Beccaccioli M, Fanelli C, Dall'Asta C, Battilani P, Reverberi M, Sanseverino W. Careful with that axe, gene, genome perturbation after a PEG-mediated protoplast transformation inFusarium verticillioides. Toxins. 2017;9(6).https://doi.org/10.3390/toxins9060183.
- 50.
Li X, Yang RH, Ma MG, Wang X, Tang J, Zhao XX, Zhang XP. A novel aldehyde reductase encoded byYML131WfromSaccharomyces cerevisiaeconfers tolerance to furfural derived from lignocellulosic biomass conversion. Bioenerg Res. 2015;8(1):119–29.
- 51.
Zhou Q, Yu BJ. Accumulation of inorganic and organic osmolytes and their role in osmotic adjustment in NaCl−stressed vetiver grass seedlings. Russ J Plant Physl. 2009;56(5):678–85.
- 52.
Wei PP, Chen DM, Jing RA, Zhao CR, Yu BJ. Ameliorative effects of foliar methanol spraying on salt injury to soybean seedlings differing in salt tolerance. Plant Growth Regul. 2015;75(1):133–41.
Acknowledgements
We thank Ms. Jee Yan Chu for copy-editing the manuscript.
Funding
This work was supported by grants from the National Natural Science Foundation of China (31671604, U1603111) awarded to BJY, and the Hong Kong Research Grants Council Area of Excellence Scheme (AoE/M-403/16), CUHK VC Discretionary Fund VCF2014004, and the Lo Kwee-Seong Biomedical Research Fund awarded to H-ML. Funding body had no role in the design of the study and collection, analysis, interpretation of data and in writing the manuscript.
Availability of data and materials
The datasets used and during the current study available from the corresponding author on reasonable request.
Author information
Affiliations
Contributions
PW, BC, LS, YC, SW, and FL conducted the experiments, collected and analyzed all data. BY and H-ML designed the experiments. BY, PW, CC, M-WL, and H-ML interpreted the data and wrote the manuscript. All authors read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Additional file
Additional file 1:
Figure S1.Pearson correlation of membrane currents ofXenopusoocytes injected expressingGsCLC-c2.Figure S2.Comparison of chloride transporting activity of GmCLC1 and GsCLC-c2.Table S1.Summary of soybeanCLC-homologous genes information inG. max(cultivar N23674) andG. soja(accession BB52).Table S2.Primers for soybeanCLCgenes analyses.Table S3.Information for site-directed mutagenesis ofGsCLC-c2 (DOCX 140 kb)
Rights and permissions
Open AccessThis article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/)适用于汽车列车的数据可用le, unless otherwise stated.
About this article
Cite this article
Wei, P., Che, B., Shen, L.et al.Identification and functional characterization of the chloride channel gene,GsCLC-c2from wild soybean.BMC Plant Biol19,121 (2019). https://doi.org/10.1186/s12870-019-1732-z
Received:
Accepted:
Published:
Keywords
- Chloride
- Chloride channels
- CLCs
- Differential expression
- GsCLC-c2
- Nitrate
- Salt stress
- Soybean
- Wild soybean